Updated: 22nd Oct 2019
Key Points
- Three species, V. cholerae, V. parahaemolyticus and V. vulnificus, are important human pathogens and are potential contaminants in seafood.
- Rapid kits now give one-day completion of enrichment and detection of pathogenic Vibrio species
- Vibrio parahaemolyticus is the most common source of seafood-borne disease but Vibrio vulnifucs has been the most lethal cause.
Members of the genus Vibrio are all Gram-negative straight or curved rods, which do not form spores and are motile, usually by a single polar flagellum. Three species are considered to be important human pathogens - V. cholerae, V. parahaemolyticus and V. vulnificus. All three have the potential to be foodborne, and are most often associated with the consumption of raw, or undercooked, shellfish.
A number of other species have infrequently been isolated from the stools of people suffering from gastroenteritis and are considered to be occasional human pathogens. These include V. alginolyticus, V. fluvialis, V. furnissii, V. hollisae, V. metschnikovii and V. mimicus. These species are not generally regarded as significant foodborne pathogens.
All of the pathogenic species have been shown to withstand freezing temperatures whilst inhabiting seafood, which is of significant concern for the seafood industry. They are also able to tolerate high pH and salinity; factors which comprise the make-up of enrichment media when isolating Vibrio sp. from other natural flora. Most Vibrio spp. are oxidase and catalase positive and ferment glucose without gas production.
They occur naturally in freshwater and marine environments and are considered the natural microflora of these environments. However pathogenic strains of these species do arise either through a rise in temperatures or through polluted waters. Many of the pathogenic species, with the notable exception of Vibrio cholerae, are adapted to salt or brackish water habitats and are halophilic to some degree, being unable to grow in the absence of sodium chloride.
Vibrio cholerae
V. cholerae is the cause of outbreaks and epidemics of cholera, a serious and potentially fatal gastrointestinal infection, which is still a major health problem in parts of the developing world. Humans are the principal reservoir for V. cholerae and cholera is usually associated with poor hygiene and polluted water supplies, but it may also be foodborne.
Contamination of fruits, vegetables and other foods usually occurs via an infected food handler, or by the use of polluted water in food preparation. Not all strains of V. cholerae cause cholera and the main virulence factor involved in the disease is the cholera enterotoxin (CT). Strains causing classic epidemic cholera generally belong to one of two serogroups, O1 or O139, although other strains have been reported to cause occasional isolated cases of cholera-like disease.
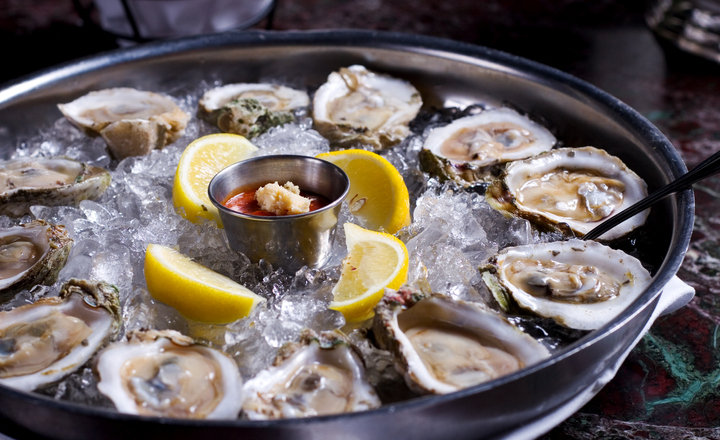
Non-O1/O139 strains can cause a less severe form of diarrhoeal disease and are more common clinical isolates in developed countries. These isolates are typically related to consumption of contaminated shellfish, especially raw oysters. Non-O1/O139 V. cholerae strains are not uncommon in certain estuarine waters and may contaminate shellfish harvested from polluted sites.
V. cholerae will grow rapidly in temperature-abused processed foods and will also survive for extended periods in chilled and even frozen foods, but it does not survive desiccation for more than 48 hours. The cells are not heat-resistant and are readily destroyed by cooking and pasteurisation.
Vibrio parahaemolyticus
V. parahaemolyticus is the species most likely to be associated with foodborne disease in humans. It can cause mild to moderate gastrointestinal infections, which are usually self limiting and rarely fatal. It is not normally involved in epidemics or large outbreaks, but is an important source of foodborne disease, especially in Japan and other Asian countries.
Infection is almost always associated with consumption of seafood and V. parahaemolyticus is very common in marine coastal waters in tropical regions and in temperate regions during the warmer months of the year. There is also evidence that it may be becoming more common in colder waters during the summer, possibly as a consequence of rising sea temperatures. As with V. cholerae, not all strains of V. paraheamolyticus are capable of causing human disease. Pathogenicity is associated with a thermostable haemolysin, called the Kanagawa phenomenon. Almost all isolates from cases of food poisoning are Kanagawa positive strains.
V. paraheamolyticus is part of the normal microflora of coastal and estuarine waters in almost all temperate regions and may be present in comparatively high numbers when the water temperature is at its highest during the summer. At these times, shellfish may easily become contaminated, but fortunately most environmental isolates are Kanagawa negative and do not cause illness. The possibility of infection means that shellfish from high-risk waters should not be eaten raw.
Most cases of V. paraheamolyticus infection in Europe can be traced to imported seafood, but there is evidence to suggest that it is becoming more common in domestically harvested shellfish. V. paraheamolyticus differs from V.cholerae in that it is an obligate halophile and will not grow unless a salt concentration of at least 0.5% is present. In fact it will grow at salt levels as high as 10%. Like V. cholerae it will grow rapidly in temperature abused foods and survives chilled and frozen storage, but not drying or mild heat processes.
Vibrio vulnificus
V. vulnificus is the leading cause of death from a seafood-borne disease in the United States. Foodborne infections can take the form of gastroenteritis in healthy adults, but in vulnerable individuals the pathogen can cause primary septicaemia, which is very serious and has a mortality rate of more than 50%. Consumption of raw oysters is the primary source of this pathogen with a significant amount of incidents related to oysters from the Gulf of Mexico that are eaten raw. The state of California has implemented a law prohibtiing the sale of raw oysters from this region applicable to certain times of the year. Only about 90 cases of V. vulnificus infection are reported each year in the USA and major outbreaks have not been recorded, but the high death rate means that it is regarded as a serious pathogen.
Elsewhere, sporadic cases have been identified in Europe, Korea and Taiwan. Like V. parahaemolyticus, V. vulnificus is a common part of the natural microflora of coastal waters in tropical and temperate regions worldwide, especially during the summer months. This being so, given the low number of cases of human infection, it is likely that only certain strains are pathogenic to man. V. vulnificus is a halophile requiring at least 0.5% salt to grow and will tolerate levels of up to 5%. It will multiply in live oysters, but not at temperatures of less than 13oC. Like other species it resists low temperatures for some time, but is destroyed by cooking and is not resistant to desiccation.
Vibrio mimicus
V.mimicus is a foodborne pathogen and has been associated with diarrhea following the consumption of raw or undercooked seafood, primarily oysters. It was isolated from samples during a search for V. cholerae as they are both nonhalophilic. V. mimicus can be differentiated from its closely related pathogen by sucrose nonfermentation.
There is currently no standards set in the revised edition of ISO 21872: 2017 for the detection of V.mimicus as was in the original ISO 21872:2007 paper. The Microbiological Methods and Bacteriological Analytical Manual (BAM), under the authority of U.S Food and Drug Administration maintains a section dedicated to the detection of V. mimicus and so worthy of mention. The organism will appear as green colonies on thiosulfate citrate bile salts sucrose (TCBS) agar and will grow in most common media without added NaCl. Virulence is poorly characterized, but some strains have been found to possess the cholera toxin gene, produce demonstrable CT in a tissue culture assay, and the ctx gene can be detected by PCR amplification.
Methods
Storage and preparation of samples The majority of food samples examined for the presence of Vibrio species will be shellfish and other seafood. It is very unusual for other food categories to be routinely tested for these pathogens
Vibrio spp can grow very rapidly in seafood at ambient temperature and samples must be chilled to below 10oC immediately and then analysed as quickly as possible. However, the cells are easily damaged by rapid cooling and samples should not be cooled by direct contact with ice.
Sample preparation procedures for shellfish typically require pooling 10-12 individual animals. The pooled sample is then homogenised using a sterile high-speed blender. If sample dilutions are required they should be prepared with a diluent containing salt, such as phosphate buffered saline (PBS).
Traditional methods
The recently revised ISO standard for the horizontal method of detection of enteropathogenic Vibrio species in the food chain now includes V.vulnificus (Standard ISO 21872-1:2017 – Microbiology of food and animal feeding stuffs — Horizontal method for the detection of potentially enteropathogenic Vibrio spp). V.vulnificus was included in Part 2 of the original document and has now been included in the revised edition due to its high rate of death after infection.
The first stage in traditional detection methods exploits the ability of Vibrio spp to grow rapidly at relatively high pH values. Media containing sodium chloride and with a pH of about 8.6, such as alkaline saline peptone water (ASPW), are used for enrichment. Typically, a 6-hour preliminary enrichment (at 41.5oC for fresh products, or 37oC for frozen or salted products) is followed by a second enrichment in ASPW at 41.5oC (for V. cholerae and V. parahaemolyticus) or 37oC (for other species) for 18 hours.
The second enrichment culture is inoculated on to thiosulphate citrate bile salts sucrose (TCBS) agar and one other optional selective medium and incubated at 37oC for 24 hours. On TCBS agar, V. cholerae colonies are smooth and yellow, V. parahaemolyticus colonies appear blue-green and V. vulnificus colonies are green or yellow. Selective chromogenic agar media specifically designed for the differentiation of pathogenic Vibrio spp are also available. Examples include chromID™ Vibrio agar from bioMérieux and CHROMagar™ Vibrio.
A quantitative method for V. parahaemolyticus can be used for samples where significant numbers are expected. This applies a MPN technique based on enrichment of tenfold dilutions in ASPW, followed by plating onto selective agar.
A hydrophobic grid membrane filtration enumeration procedure (HGMF) has also been described.
Rapid methods
There are more rapid PCR based test kits for Vibrio spp. in foods becoming commercially available than was previously. It is now possible to complete both the enrichment and detection stage in one day. An example of this is the Bio-Rad iQ-Check Vibrio Real-Time PCR Detection Kit which was released onto the market last year. It can enrich and detect the three main pathogenic species of Vibrio in just eleven hours.
Another example of a commercially available PCR-based method for pathogenic Vibrio detection is the BAX® System Real-Time PCR Assay from Hygiena. Part of the well established family of BAX® System assays, the Vibrio assay is able to detect the three most important species, V. cholerae, V. parahaemolyticus and V. vulnificus in the same sample within 24 hours, including an 18-20 hour enrichment. It is designed for testing seafood samples, including shrimp, oysters, crabs and tuna and is capable of detecting 104 CFU/ml.
Confirmation and identification
Traditional methods There are well established confirmation and identification procedures for pathogenic Vibrio spp., especially for V. cholerae. Preliminary identification based on colony appearance on TCBS agar is traditionally confirmed using classical biochemical tests. Key tests are oxidase reaction and the presence or absence of lysine and ornithine decarboxylases and arginine dehydrolase. Media should be prepared with 2-3% sodium chloride to allow the growth of halophilic species.
V. cholerae isolates are further confirmed and characterised by serological agglutination testing, β haemolysis on blood agar and tests for cholera toxin production.
V. parahaemolyticus isolates can also be characterised by serological testing and by the Kanagawa test.
Rapid methods
As an alternative to the gold standard method of using TCBS agar, Hardy diagnostics have the HardyCHROM™ Vibrio, a one-plate-for-all that allows the differentiation between all species. Pathogenic species show up different colours from each other with the aid of fluorescent light and non-pathogenic species show no colour.
Biochemical confirmation can be accomplished using commercial identification systems such as the API 20E test strip from bioMérieux and Remel’s RapID™ NF PLUS System. However it is important to ensure that cultures are suspended in a saline medium to ensure the growth of halophilic species.
A number of molecular methods for confirmation have been developed, notably PCR assays for the identification of the three most important species. Labelled DNA probes can be used to confirm the pathogenicity of V. parahaemolyticus and V. vulnificus by detecting the genes for specific virulence factors, such as haemolysins, and a PCR method has been developed for cholera toxin, which is rarely present in V. cholerae isolates from food. Protocols for several of these methods are provided in Chapter 9 of the USFDA BAM and the revised Standard ISO 21872-1:2017 – Microbiology of food and animal feeding stuffs — Horizontal method for the detection of potentially enteropathogenic Vibrio spp
Get the latest updates in Rapid Microbiological Test Methods sent to your email? Subscribe to the free rapidmicrobiology eNewsletter