Clever Culture Systems’ Automated Plate Assessment System, APAS®, brings intelligent automation to microbiology. The APAS Independence fully automates the interpretation of bacterial growth on culture plates to quickly identify and separate those plates with colonies requiring further investigation – freeing up microbiologist time to focus on more complicated cases.
Clever Culture Systems’ Automated Plate Assessment System, APAS®, brings intelligent automation to microbiology. The APAS Independence fully automates the interpretation of bacterial growth on culture plates to quickly identify and separate those plates with colonies requiring further investigation – freeing up microbiologist time to focus on more complicated cases.
Using sophisticated algorithms and a microbiology decision support system that views culture plates in the same way as a microbiologist;
APAS is able to read and interpret plates in as few as 6 seconds without human intervention.
APAS efficiently sorts the plates into three categories: positive, negative or no significant growth, and those requiring immediate review, such as those with multiple colony types.
With negative plates removed from the work stream, microbiologists can quickly attend to and focus on significant cases. The consequent delivery of faster results reduces laboratory reporting times.
APAS will improve your work-flow which in turn, improves the timeliness of result delivery; saving time and costs, but more importantly, contributing to saving lives.
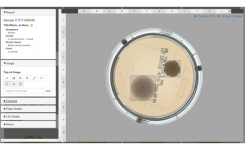
APAS automatically produces informative reports to enable the swift delivery of critical results for optimal patient management. These reports are designed by clinical microbiologists for clinical microbiologists to ensure relevance and usefulness.
- Categorizes colonies into well recognised morphological groups
- Combines results for multiple plates for each sample
- Sample specific reporting
- Incorporates additional clinical data such as urine white cell count
- Based on conventional microbiological reporting rules including:
- Agar plate manufacturers’ recommendations
- Expert rule sets and published guidelines (USA, European, UK etc.)
- High quality image review capability
- Workload reporting to assist with laboratory workload management.