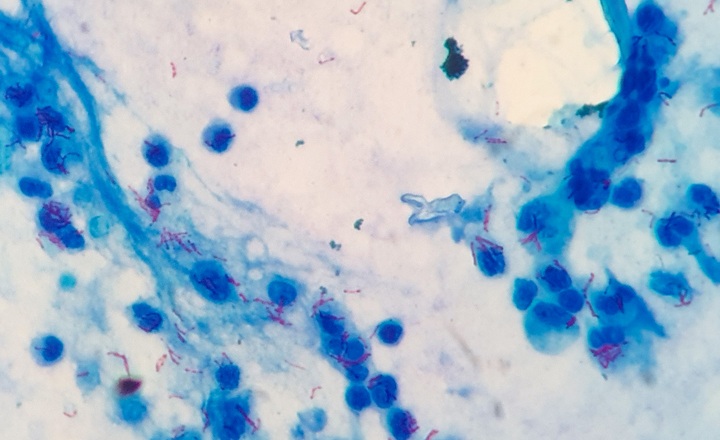
- Slow growth: Many mycobacterial species, including Mycobacterium tuberculosis, require weeks to grow in culture, delaying diagnosis and treatment decisions.
- Limitations of culture: Some species grow poorly or may be outcompeted by contaminants, leading to under-detection of clinically relevant mycobacteria.
- Need for comprehensive identification: Accurate species and subspecies identification is essential because treatment regimens vary across Mycobacterium species.
- Potential of culture-free sequencing: Target capture sequencing can detect mycobacterial DNA directly from specimens, potentially accelerating diagnosis and expanding detection of rare or fastidious species.
Key Findings: Hashimoto et al. undertook a prospective comparison of conventional culture workflows and a culture-free target capture sequencing method that identifies mycobacteria directly from sputum samples.¹ The authors analyzed 125 sputum specimens from 115 patients with nontuberculous mycobacterial (NTM) pulmonary disease and 10 with non-NTM respiratory conditions.
- Detection performance: Method accuracy depended strongly on bacterial load, reaching 75.4% in smear-positive specimens but only 13.9% in smear-negative specimens.
- Subspecies resolution: Target capture sequencing enabled subspecies identification using cgMLST, including Mycobacterium avium subsp. hominissuis (48), Mycobacterium intracellulare subsp. intracellulare (22), subsp. chimaera (5), Mycobacterium abscessus subsp. abscessus (7), subsp. massiliense (5), and one M. tuberculosis isolate. Specific NALC-NaOH pretreatment enabled subspecies-level identification with > 90% accuracy.
- Workflow advantage: Because sequencing is performed directly on clinical samples, results can potentially be generated within a single day, avoiding the prolonged incubation required for mycobacterial culture.
Bigger Picture: Culture remains the clinical gold standard for diagnosing mycobacterial infections as it provides viable organisms for antimicrobial susceptibility testing. However, the slow growth and limited recovery of some species constrain its diagnostic utility. This study demonstrates that culture-free sequencing approaches can substantially expand the detection and characterization of mycobacteria directly from patient samples, potentially improving diagnostic speed and taxonomic resolution. In the future, combining culture-independent sequencing with traditional culture workflows may provide the most comprehensive strategy—rapid identification from sequencing followed by culture for susceptibility testing and epidemiologic analysis.
(Image Credit: iStock/ DouglasOlivares)