Frequently Asked Questions on Water Testing
- What are indicator organisms? Indicator organisms such as coliforms are associated with faecal contamination and are always present when enteric pathogens and viruses are present.
- Why are indicator organisms used in water testing? Indicator organisms are used as they are an easy-to-detect group rather than having to test for each specific pathogen; their presence is an 'indicator' of faecal contamination
- Are all coliforms indicator organisms? No, many coliforms are found in plants and soils they are not necessarily of faecal origin. where coliforms are detected a second step is needed to identify faecal coliforms from coliforms found in the environment.
- What are the names of the faecal coliforms? Most common ones are Escherichia, Enterobacter, Klebsiella are the faecal coliforms, whilst Citrobacter and Serratia are often found in plants and soil.
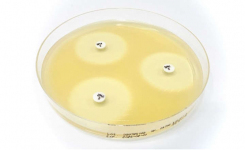
- Why is membrane filtration used for water testing? Membrane filtration allows a larger volume of water to be tested than would otherwise be possible. The water sample is filtered through a membrane normally with a pore size of 0.45µm, microorganisms are caught on the membrane which is then laid onto microbiological growth media and incubated, to allow for colonies to grow and be counted.
Programme of Testing
The most effective way to check water supplies for faecal contamination is microbiological analysis, and a range of test methods designed for that purpose has been developed for the water industry. Instead of carrying out separate tests for each of the potential pathogens, viruses, or parasites that might be in the water, microbiologists test for indicator organisms that are always present when enteric pathogens and viruses are.
Coliforms and Other Indicator Organisms
Coliform is a term used to denote a group of Gram-negative bacteria that can ferment lactose with a production of gas within 48 hours at either 35ºC or 44/44.5ºC. These characteristics allow for easy isolation, detection, and enumeration in the lab and are the gold standard for microbial water testing. They are always present when enteric pathogens or viruses are detected in water testing. However, a high 'total coliform' count doesn't necessarily mean faecal contamination and requires a second step to identify the faecal coliforms from coliforms found in the environment. Escherichia, Enterobacter, Klebsiella are the faecal coliforms, and Citrobacter and Serratia are found in plants and soil.
The faecal coliforms, also known as thermotolerant coliforms, can survive at temperatures of 44ºC and 44.5ºC, which allows simple differentiation between the two types. The microbiologist will be looking for counts of faecal coliforms such as E. coli, whose only habitat is the intestine, and whose life outside it is short-lived, and is seen as the ideal indicator organism. Its presence in a sample of drinking water means that the water is unsafe for human consumption.
The presence of faecal streptococci/Enterococci is evidence of faecal contamination. Faecal streptococci tend to persist longer in the environment than thermotolerant or total coliforms and are highly resistant to drying. It is, therefore, possible to isolate faecal streptococci from water that contains few or no thermotolerant coliforms as, for example, when the source of contamination is distant in either time or space from the sampling point. Faecal streptococci grow in or on a medium containing sodium azide, at a temperature of 37-44 °C. They are usually detected by the reduction of a dye (generally a tetrazolium-containing compound) or the hydrolysis of aesculin. Conventional methods can give 'false positives' and additional confirmatory tests may be required.
Rapid Methods
Defined substrate technology® (DST) developed by IDEXX can produce results in 24 hours. The IDEXX Colilert uses a colourimetric ONPG assay to detect coliforms and a fluorescence MUG assay for E. coli. Colilert can simultaneously detect these bacteria within 18-24 hours, depending on the product. It can also suppress 2 million heterotrophic bacteria per 100 mL present. It has been U.S E.P.A approved and is included in the U.S Standard Methods for Examination of Water and Wastewater. As of 2014, this technology from IDEXX has been published as a European Standard Method, and many countries now use this technology as their gold standard for water testing e.g. Finland and Ireland.
The Enterolert Test from IDEXX uses a proprietary Defined Substrate Technology (DST) nutrient indicator to detect enterococci. This nutrient indicator fluoresces when metabolized by enterococci. DST improves accuracy and avoids the need for hazardous sodium azide suppressants used in traditional media.
Membrane Filtration
A typical MF method for water analysis is performed by passing a known volume of water through a sterile membrane filter with a pore size small enough to retain bacterial cells (typically 0.45µm). The filter is then transferred aseptically to the surface of an agar plate, or an absorbent pad saturated with a suitable selective medium and incubated. Colonies are allowed to develop on the surface of the filter and can be counted and examined directly. MF methods are quick and easy to perform, require little incubator space and can handle very large volumes of water if required.
Over the last 30 years, they have become the preferred methods for the microbiological examination of water for indicator organisms. There are several official published methods based on MF, notably a series of ISO methods, such as ISO 9308-1 for coliforms and E. coli and ISO 7899-2 for enterococci.
The US Environmental Protection Agency (EPA) has published official microbiological methods for water testing. Laboratories routinely testing drinking waters, recreational waters and environmental samples should use the appropriate official method recommended by their local enforcement agency. Laboratories testing water supplies for industrial use, such as food processing, are advised to use the same methods when the water supply is required to be of potable quality.
In terms of equipment, MF methods require suitable filtration apparatus consisting of a base supporting a porous disc, on which the filter is placed, and a sterile filter funnel, which can be secured to the base, clamping the filter in position. A variety of filtration systems are commercially available, including multiple units in the form of manifolds, so that more than one sample can be filtered at once.
Disposable single-use sterile filter funnels are also available for convenience. Suppliers include Millipore, Sartorius and Membrane Solutions. The filter unit is connected to a suitable vacuum source to draw the samples through the filter. This may be a vacuum line or a stand-alone pump unit. Compact pump units specifically designed for use with MF methods are now available. An example is the Millipore EZ-Stream unit, which can run the filtered sample straight to drain, thus saving time spent emptying and cleaning the waste sample containers used with traditional laboratory vacuum pumps. A variety of membrane filters are available for different applications, but microbiological water analysis is typically carried out with 47mm diameter mixed cellulose ester-based filters of 0.45µm pore size. The filters are usually marked with a grid to aid colony counting.
Culture Media
Much selective media have been developed for the detection of indicator organisms in water by MF methods. Recommended media for coliforms and E. coli include membrane lauryl sulphate broth or agar, MI agar and broth, and membrane lactose glucuronide agar. Membrane enterococcus agar (mEA) and membrane-Enterococcus Indoxyl-ß-D-Glucoside Agar (mEI) can be used for detection and enumeration of enterococci, while Tryptose sulphite cycloserine agar without egg yolk can be used to culture Clostridium perfringens on membrane filters. Pseudomonas aeruginosa can also be detected by an MF method using Pseudomonas agar.
Further culturing, or biochemical testing can then be used to confirm the identity of suspect colonies growing on filters placed on selective media. Increasingly, chromogenic and fluorogenic media are being used in water microbiology. Based on the detection of specific enzymes in the target bacterial species by substrates containing chromogenic or fluorogenic groups, producing highly diagnostic coloured colonies, these media can be less harsh than other selective media, resulting in fewer false-negative results, and reduce the time needed to confirm results. Examples include the ChromoCult® media range produced by Merck and the BBL™ range of prepared chromogenic media for water testing.
Traditional Culture
Techniques using pour and spread plate count methods are not sufficiently sensitive for the detection of indicator organisms and pathogens in water, although they are still used routinely for enumerating heterotrophic bacteria. Methods capable of testing a larger volume of water (typically 100 ml) are needed. For many years the method of choice was the multiple tube ‘most probable number’ (MPN) technique, in which measured volumes of the water sample are added to a series of tubes containing differential media and incubated.
Growth is indicated by a colour change in the medium, and the result is calculated from the distribution of positive tubes. Although the method is simple and inexpensive in terms of equipment and materials, it is labour intensive and requires large amounts of incubator space. It is also an indirect method and does not allow further examination of individual colonies. MPN tests for routine water microbiology have now been largely replaced by membrane filtration (MF) methods, although they may still be useful for occasional tests conducted in small laboratories or the field, and commercial test kits based on MPN methods are available for coliforms and enterococci.
Legionella in Cooling Towers
Testing for Legionellasp. in cooling towers should be carried out regularly and must be part of a plant's programme of testing. Legionella pneumophila, for instance, is a heat-loving organism that has a higher tolerance for chlorine than most bacteria and can live within the cells of parasites, protecting itself from these methods of cleaning. Cooling towers provide suitable conditions for L. pnuemophila and unfortunately provide a perfect transport for its dissemination via droplets into the atmosphere whereupon it is potentially inhaled by people in the vicinity and cause pneumoniae. For information on products for testing water for Legionella go to: Legionella Detection and Identification Methods
Water Quality Parameters
In addition to tests for indicator organisms and certain specific pathogens, non-selective colony counts are also routinely carried out to determine the population of heterotrophic bacteria present. Counts at two temperatures (22oC and 37oC) are typically performed to provide information on the general microbiological population of the water and detect sudden changes in water quality. Counts at 37oC have been used to indicate faecal contamination in the past, but this is not generally considered to be reliable.
Rapid Methods
Although most official methods for microbiological water analysis still rely on traditional culture methods and MF methods, the time is taken to obtain results, typically 24-48 hours, has focused attention on alternative rapid methods. Combining MF with QPCR detection and enumeration is a particularly rapid and effective means of analysing water samples. The main disadvantage of this method is that it may detect non-viable cells and overestimate the population, but it seems likely that QPCR-based methods will become increasingly important in water microbiology, leading to the development of commercial products similar to those already used for food analysis.
During the SARS-CoV-2 outbreak, traces of the viral RNA was found in sewage at treatment plants in one study carried out in The Netherlands. The virus was detected by the use of RT-PCR against three fragments of the nucleocapsid protein gene (N1-3) and one fragment of the envelope protein gene (E), with the N1 primer/probe set being the most sensitive, followed by N3 and E sets for detection of SARS-CoV-2 in sewage.
In recent years, water microbiologists have become increasingly aware of the importance of biofilms for microbiological populations in water systems. Biofilms are now recognised as complex microbial communities, which form on surfaces. Biofilms typically consist of a variety of microbial cells, potentially including pathogens, within a matrix composed of exopolysaccharides (EPS) secreted by certain bacterial species. Biofilms develop over time, becoming more complex and extensive, and can protect individual bacterial cells from chlorine and other antimicrobial compounds in water.
Biofilms are also notoriously difficult to remove from surfaces and can act as a sporadic source of microbial contamination as bacterial cells are sloughed off from the matrix into the surrounding water. It is now recognised that most of the bacteria in drinking water distribution systems are present within biofilms rather than free-living in the water itself. Pathogens isolated from within biofilms include Salmonella Typhimurium, Campylobacter sp. Pseudomonas aeruginosa and Aeromonas hydrophila. Legionella pneumophila is now known to be a major pathogen in residential water systems, due to its proclivity to form biofilms on water distribution pipes and also within cells of microbial parasites. Biofilms may affect general microbiological water quality, cause objectionable tastes and odours and accelerate corrosion within distribution systems.
The presence of significant biofilm growth may make it difficult to obtain representative water samples and may influence the results of microbiological analysis. High heterotrophic plate counts may be indicative of biofilm formation in distribution systems. In some cases, it may be necessary to sample biofilms directly using swabs or by allowing a film to develop on the surface of removable metal coupons or within specially designed sections of pipework.