Key Points
- Traditional colony count methods for yeasts and moulds require 5-7 days to achieve a result. Rapid detection methods can reduce this by 50% or more.
- The interpretation of conventional colony counts can be difficult, requiring considerable staff input. Rapid methods can help reduce the possibility of human error and free up skilled technicians for other tasks.
- Automated rapid detection methods have the potential to provide significant time and cost savings for some laboratories.
Until about 50 years ago the spoilage of food by microorganisms was regarded as a quality issue rather than a food safety concern. Since then, the discovery that many common mould species are in fact dangerously toxigenic has led to a radical re-think. Mycotoxin production in foods is now recognised as an important threat to public health and considerable attention is given to its control. Nevertheless, fungal spoilage of food remains a serious problem, one that has been estimated to account for between 5% and 10% of all losses in global food production. Despite this, methods for the detection, and especially the identification, of food spoilage yeasts and moulds have undergone relatively little development in comparison with methods for bacteria. Recently this situation has begun to change and some of the technological advances developed for the detection of pathogenic and spoilage bacteria have been adapted for rapid detection of yeasts and moulds.
Food spoilage by microfungi
Hundreds of different species of yeasts and moulds have been isolated from foods and their ability to grow over a wide spectrum of environmental conditions mean that very few foods are entirely safe from fungal spoilage. Yeasts and moulds are able to grow over a wide range of temperatures and pH values, but it is also the ability of many species to grow at reduced water activity (aw) values that equips them to be such important agents of food spoilage. Some important food spoilage moulds, such as Eurotium species, Wallemia sebi and certain Aspergillus and Penicillium species, are able to grow slowly at aw values down to about 0.70, while the most xerophilic organism known, Xeromyces bisporus, has been shown to grow at wter activity (aw) 0.61 and has been found in a variety of low moisture foods.
An enormous range of different foods can be spoiled by yeasts and moulds, including fresh fruits and vegetables, nuts, beans and pulses, cereals, meat and dairy products, flour, pasta, bakery products, jams and preserves, soft drinks and alcoholic beverages, confectionery, canned acid foods, cheeses and processed meats. Spoilage may be highly visible, in the form of surface fungal mycelium or obvious degradation of the product, or may be less evident and only detectable as an ‘off’ flavour, or fermentation and gas production.
|
Some Important Food Spoilage Yeast and Mould Species |
|
|
MOULDS
|
|
|
Species
|
Foods affected
|
| Aspergillus flavus | Nuts and oilseeds (mycotoxin producer – aflatoxins) |
| Aspergillus niger | Fresh fruit and vegetables, nuts and seeds, dried foods |
| Aspergillus ochraceus | Dried and stored foods (mycotoxin producer – ochratoxin) |
| Byssoclamys fulva | Canned fruits |
| Eurotium chevalieri | Cereals and a wide range of processed and stored foods |
| Fusarium culmorum | Infects cereals, especially barley, in the field (mycotoxin producer – DON and others) |
| Penicillium aurantiogriseum | Stored fruits and vegetables |
| Penicillium commune | Cheeses |
| Penicillium digitatum | Citrus fruits |
| Rhizopus stolonifer | Fruits, tomatoes and peppers |
| Wallemia sebi | Dried and salted fish and other dried foods |
|
YEASTS
|
|
| Brettanomyces bruxellensis | Beer and wine, fruit yoghurts |
| Debaryomyces hansenii | Cured meats and brined products |
| Saccharomyces cerevisiae | Soft drinks and fruit juices |
| Zygosaccharomyces bailii | Soft drinks, sauces, fruit juice, wine, ciders and syrups |
| Zygosaccharomyces rouxii | Confectionery, fruit concentrates |
Traditional methods
Storage and preparation of samples
A very wide range of foods and beverages may be routinely tested for the presence of yeasts and moulds. Both fresh and processed products are subject to fungal spoilage and even heat-processed foods may be affected. Environmental samples may also be tested on occasion, particularly from surfaces in storage areas, chillers and processing lines.
Food and environmental samples should be tested as soon as possible after collection. If this is not possible, samples should be stored at 4oC, but should not be frozen.
Unless the sample is a liquid, an initial suspension is prepared by mixing the sample with a suitable diluent such as peptone water broth. It may be beneficial to add a small quantity of a suitable surfactant to the diluent (e.g. polysorbitan 80) to help break up clumps of mould spores and conidia. When testing dried products and concentrates for xerophilic moulds and yeasts, 20-30% sucrose or glycerol can be added to the diluent to prevent osmotic shock. Very dry, hard foods, such as pulses, should be allowed to soak in the diluent for up to 60 minutes before homogenisation.
Detection and enumeration
Most routine tests for yeasts and moulds in food samples use conventional colony count methods rather than specific detection techniques.
There are current ISO horizontal methods for the enumeration of yeasts and moulds in foods and animal feed. ISO 21527:2008 is published in two parts. Part 1 relates to food and feed samples with water activity of >0.95, while part 2 applies specifically to dried and processed foods with reduced water activity of <0.95. A further standard, ISO 6611:2004, is a method designed for the enumeration and yeasts and/or moulds in milk and milk products. Similar standard methods have been published elsewhere by other bodies, notably in the USFDA Bacteriological Analytical Manual (BAM).
These methods typically employ a surface plating technique, where a known quantity of the sample, or the initial suspension, is spread over the surface of a suitable selective agar medium. ISO 21527-1 recommends Dichloran Rose Bengal Chloramphenicol agar (DRBC), while the preferred medium specified in 21527-2 is Dichloran 18% Glycerol agar (DG18). Other media used include Oxytetracycline Glucose Yeast Extract agar (OGYE) and Yeast Extract Dextrose Chloramphenicol agar (YGC). Selective agars for yeasts and moulds usually contain antibiotics to help suppress bacterial growth. Plates are typically incubated at 25oC for 5 to 7 days and then examined for the presence of yeast and mould colonies.
Isolation and identification
Individual yeast and mould species can be isolated from selective agars by subculture onto a non-selective agar such as Malt Extract agar (MEA) or Potato Dextrose agar (PDA). Although it may be possible to recognise some moulds at the genus level simply by colony morphology and the appearance of conidia and other features under the microscope, identification to species level is very difficult and requires specialised skills and experience. Identification of yeast and mould species is therefore best left to reference laboratories and recognised experts.
Rapid methods
Detection and enumeration
The lengthy incubation times needed for conventional yeast and mould counts have driven a number of attempts to develop rapid detection and enumeration methods, especially for food spoilage yeasts. Some of the technologies employed include conductimetry, ATP assay, immunological techniques (e.g. ELISA) and molecular biology-based methods. Relatively few of these have been successfully developed into commercially available systems for the food industry, but there are now a handful of products designed for food microbiology applications.
Flow-Cytometry Chemunex technology from bioMérieux provides rapid detection of Yeast and Molds in a wide range of product types such as juices, iced teas, beverages, fermented milk products and yogurts. Objective YES/NO results are generated within minutes following incubation of 50 hours for moulds.
Growth-based methods
The majority of the rapid methods currently available for detection and enumeration of yeasts and moulds in food and drink samples are culture-based, but utilise various means to cut incubation times. Several systems rely on the replacement of Petri dishes by membrane filters, or dried culture media enclosed in plastic or fabric films. These systems often have a built-in grid to help colony counting and are claimed to cut incubation times from five days to 48-72 hours. Examples include 3M™ Petrifilm™ Yeast and Mould Count Plates.
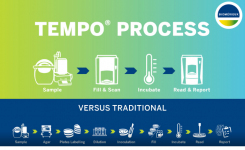
Growth detection systems have been taken a step further in instruments designed to continuously monitor liquid media inoculated with a food sample for indicators of microbial growth. These automated optical devices use pH change and other biochemical reactions occurring during microbial growth to generate visible colour changes, or fluorescence in the medium, which can be monitored photometrically. A double-chambered test vial allows the colour change to be monitored without interference from the food matrix. Two examples of such instruments are the Soleris® system from Neogen and the BioLumix system. Both have been developed for food applications and can be used to detect and enumerate yeasts and moulds within a claimed 72 hours. The BioLumix system detects optical changes in an embedded optical sensor for the detection of CO2, located at the bottom of the test vial. Yeast and mold growth in the medium above the sensor results in CO2 production and changes in the sensor’s color within 48 hours. A third growth-based instrument is the TEMPO® “automated quality indicator” system from bioMérieux. This employs a miniaturised most probable number (MPN) technique on a multi-well ‘card’ to enumerate microbial counts in food samples and can be used to enumerate yeasts and moulds. The media used in the system contain a fluorescent indicator. The MPN card is scanned by a reader, which detects any fluorescing wells and then automatically calculates a result.
Molecular methods
Although it has proved difficult to develop immunological methods that will detect a sufficiently broad spectrum of yeast and mould species, molecular biology-based methods have been successfully commercialised and automated for food industry applications. A yeast and mould assay kit has been developed for the widely used BAX® System from Dupont Qualicon, which amplifies and detects a DNA sequence specific to microfungi. The kit is able to detect yeasts and moulds at levels of 50 CFU/g within hours after a 44-hour enrichment step, but can detect higher levels within a day.
A test for fermentative yeasts designed specifically for the beverage industry has been developed by vermicon AG. Based on gene probe technology, the VIT-Fermentative Yeasts test kit is claimed to detect low levels of viable yeast cells with a time saving of several days over conventional methods.
Identification
Few rapid methods are available that can be used to identify yeast and mould isolates from food products. However, several systems have been developed to identify fungi primarily from clinical samples and these may be of use for food applications. The best known and most widely used in food microbiology labs are Biolog automated and manual systems for yeasts and filamentous fungi and the API® 20C AUX and ID 32 C strips for identifying yeasts. Both use carbon source metabolic ‘fingerprints’ to obtain a preliminary identification within 24-48 hours.
Get the latest updates in Rapid Microbiological Test Methods sent to your email? Subscribe to the free rapidmicrobiology eNewsletter