In recent years, the global prevalence of Legionnaires’ disease has steadily increased. According to the European Legionnaires’ Disease Surveillance Network’s (ELDSNet) 2017 "Surveillance and outbreak report", the ASR (age-standardised rate) for Legionella in Europe increased from 0.97 to 1.30 cases per 100 000 population, which corresponds to an annual increase of 0.09 cases per 100 000 population. Legionnaires' disease is caused by gram negative bacteria belonging to the species Legionella. These pathogens commonly occur in natural freshwater sources and can often be found in man-made water systems, such as water heaters and cooling towers. Infections occur through the inhalation of aerosols.
The confirmed and identified cases between 2011 and 2015 were predominantly caused by Legionella pneumophila, more specifically, Legionella pneumophila subsp. pneumophila serogroup 1 (SG1). People aged 50 or older were identified as the most sensitive group, with 80% of the infections falling into this age demographic.
One of the main issues regarding the prevention of Legionnaires' disease is the time to detection and identification of the pathogen in water samples. The current recognised ISO method requires an initial incubation period of 10 days for colony growth and a further 5 days for the confirmation of Legionella colonies. For identification additional methods, such as MALDI-TOF or agglutination tests, have to be implemented. However, while MALDI-TOF is fast and cost-efficient after the initial investment in the instrument, this method lacks the resolution to differentiate the most relevant serogroups and the more classical latex test fails to detect several of the relevant Legionella species other than L. pneumophila.
The BACGene Legionella Multiplex kit is a newly developed real-time PCR assay, which simultaneously confirms and identifies Legionella spp., Legionella pneumophila and Legionella pneumophila subsp. pneumophila SG1.
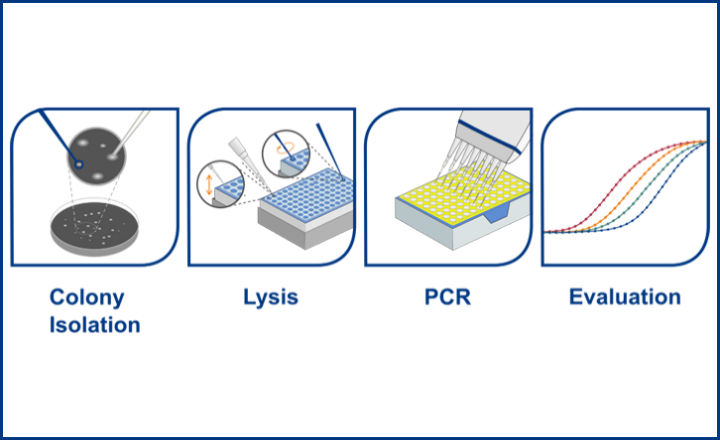
The validated workflow is a colony PCR, which can be performed from any of the initial isolation plates of the ISO method. Colonies can be picked as soon as they can be rated as presumptive positive and placed directly into the PCR after a simple thermal lysis step. This enables the confirmation and identification of presumptive positive colonies prior to the end of the initial incubation, thereby eliminating the need for a re-streaking and secondary incubation period. Therefore the BACGene Legionella Multiplex protocol leads to a significant reduction of agar plates and incubator space needed, as well as time to result. Additionally, the earlier a presumptive colony can be confirmed and identified the less likely it is to be overlooked due to overgrowth by background flora on the agar plates.
Unlike the latex identification test, which requires three separate tests, the BACGene Legionella Multiplex assay allows the identification of all three phylogenetic groups in one reaction. This reduces both handling time and space needed, due to a smaller number of plates required. In addition to this, the BACGene Legionella Multiplex kit is able to identify all relevant Legionella species as defined by the ISO 11731: 2017, some of which cannot be detected by the latex test. The BACGene Legionella Multiplex kit also identifies Legionella pneumophila subsp. pneumophila SG1, which is not identified by MALDI-TOF.
Advantages of BACGene Legionella Multiplex:
- Simultaneous confirmation & identification
- Three phylogenetic groups identified in one real-time PCR reaction
- IPC
- Identification of Legionella pneumophila subsp. pneumophila SG1
- Reliable identification of Legionella spp.
- Time, space and cost saving