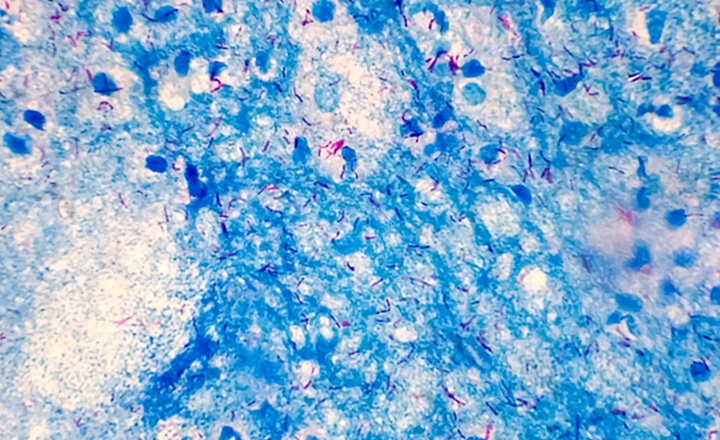
- Isoniazid (INH) is a cornerstone first line antitubercular antibiotic used for treatment worldwide. Its resistance is the most common form of drug resistance in TB and often precedes rifampin resistance in the evolution of multidrug resistant TB (MDR TB), making early detection critical for effective case management.
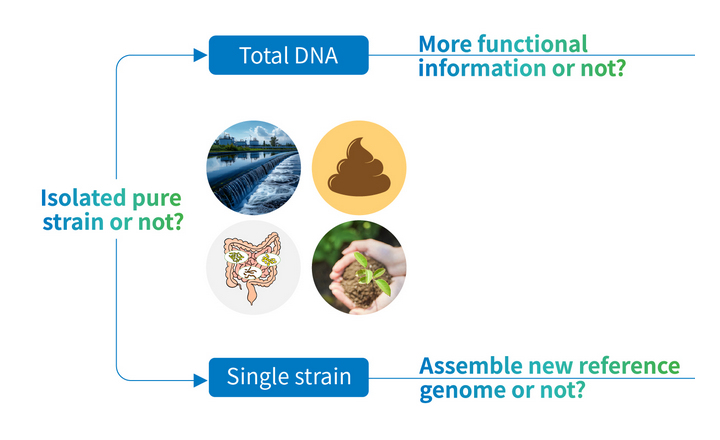
- Traditional phenotypic drug susceptibility testing (pDST) for MTB complex is slow (often weeks to months) due to the organism’s slow growth. WGS offers rapid, high resolution genomic analysis that can identify known resistance associated mutations.
- Accurate genomic prediction supports streamlined laboratory workflows, reduced turnaround time, lower costs, and earlier initiation of appropriate therapy, which is essential for patient care and public health surveillance.
Key Findings: This report by Patel et al. details a 2 phase study using WGS and phenotypic drug susceptibility testing (pDST) to assess the predictive performance of WGS for Isoniazid (INH) resistance. The study was conducted over six years and involved 3,696 isolates from New York State.1
- Prevalence and algorithm development: The overall INH resistance prevalence was 10.2%. Based on performance data, the authors developed a molecular testing algorithm that reduces reliance on pDST for isolates with clear WGS predictions, streamlining workflows and lowering costs.
- High predictive performance: In the initial comparison of 1,767 isolates with paired phenotypic and genotypic testing, WGS predicted INH resistance with 90.3% sensitivity and 99.8% specificity. The negative predictive value for susceptibility was 98.8%, indicating excellent ability to rule out resistance when no resistance mutations are found.
- Mutation basis: The majority of phenotypic INH resistance was explained by known mutations in genes such as katG, inhA, mabA, oxyR ahpC, demonstrating that most resistance has a molecular basis detectable by WGS.
Bigger Picture: This study adds to growing evidence that whole genome sequencing can serve as a rapid, accurate molecular alternative to traditional phenotypic drug susceptibility testing for tuberculosis. WGS enables early identification of drug resistance directly from cultured isolates and increasingly from primary specimens, shortening turnaround times and supporting earlier, mechanism based therapeutic decisions.
WGS also captures broader genomic context, including lineage and outbreak information, enhancing diagnostic and surveillance capacity. Implementation of WGS based predictions aligns with strategies by public health agencies and WHO to move toward genomic DST for TB, particularly in settings with established sequencing infrastructure.
Ongoing challenges include the need to continually update resistance mutation catalogs, interpret variants of uncertain significance, and maintain bioinformatics capabilities. However, this study demonstrates that, for key drugs like isoniazid, WGS can reliably predict resistance and support laboratory algorithms that reduce phenotypic testing burden while maintaining clinical accuracy.
(Image Credit: iStock/Md Ariful Islam)