Nowadays, Artificial Intelligence (AI) and machine learning are used for various applications. Microbiology has shown us a new way of deploying its power: by giving an unexpected second wind to the oldest but yet most robust way of detecting and enumerating microorganisms… culturing on agar.
How so? By combining traditional cell-marking and optical technologies, AI and machine learning make it possible to identify targeted microorganisms at their initial stage of growth: the microcolony stage.
The traditional Pasteurian detection method required a technician to wait for microorganisms to replicate and form colonies visible to the naked eye, therefore allowing enumeration. Depending on the targeted microorganism, the incubation time can vary from 24 hours to several days, and the recognition of colonies is subject to human interpretation. An identification procedure followed these detection and enumeration processes to confirm the colonies counted were the ones they were actually looking for.
But that, was before...
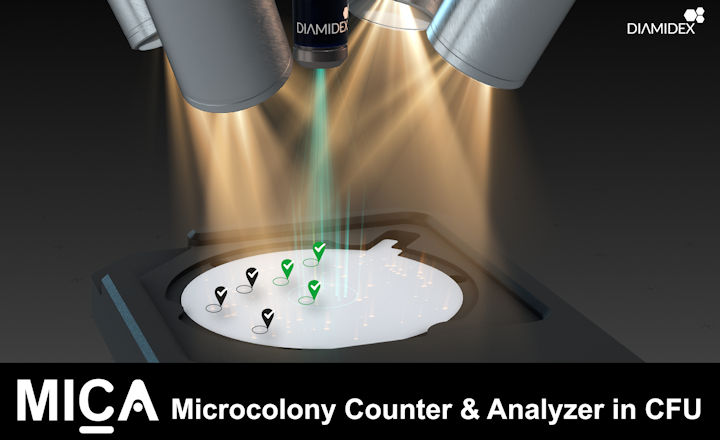
In autumn 2020 MICA, was released by DIAMIDEX, a French company specializing in the rapid enumeration of target microorganisms. MICA, which stands for Microcolony Counter and Analyzer, is a smart solution that allows to detect, identify and enumerate any microorganism at its microcolony stage present in filterable samples… In other words, much faster than the traditional Pasteurian method.
The MICA analysis procedure relies on the international standard culture methods (ISO, IFU), which implies that all strategic treatments performed on samples (acid shocks, thermal shocks, sample preparation etc.) are respected.
The technician will conduct the usual procedure, with one slight adjustment: a single drop of Diamidex marking reagent is placed on the agar plate before placing the membrane onto it. And that’s it! MICA counter automatically scans the membrane for the presence of the targeted microorganism.
MICA high-tech optical components take dozens of high-definition pictures of the membrane, allowing to detect the presence of objects measuring less than 10µm. Then, MICA AI analyses them carefully for identification, eliminating all fluorescent particles which do not correspond to the targeted microcolonies (debris, artefacts, other microorganisms, any other particle, etc.). Once recognized, the microcolonies are counted, and MICA tells you how many Colony Forming Units (CFU) of the targeted microorganism are present in your sample.
Available software:
- MICA Legionella, allowing to detect and enumerate Legionella pneumophila of all serogroups in just 48 hours (instead of 10-14 days with the ISO 11731 standard method)
- MICA Alicyclobacillus, detecting and enumerating all problematic Alicyclobacillus spp. in 24 hours, including important fruit-juice spoilage organism A. acidoterrestris (24 hours instead of 5 days using standard IFU Method No.12)
The MICA counter is compatible with both software, making it a multiparameter, smart and affordable technology.
Next in Line:
- MICA hardware and software for detection on chromogenic agar without any additional reagent: Coliforms, Salmonella spp. & Listeria spp.
- MICA software using fluorescent detection and Diamidex reagent: Total Mesophile Aerobic Flora, Yeasts & Molds, Acetic Acid Bacteria, Lactic Acid Bacteria, and so much more!
Please visit DIAMIDEX to find out more