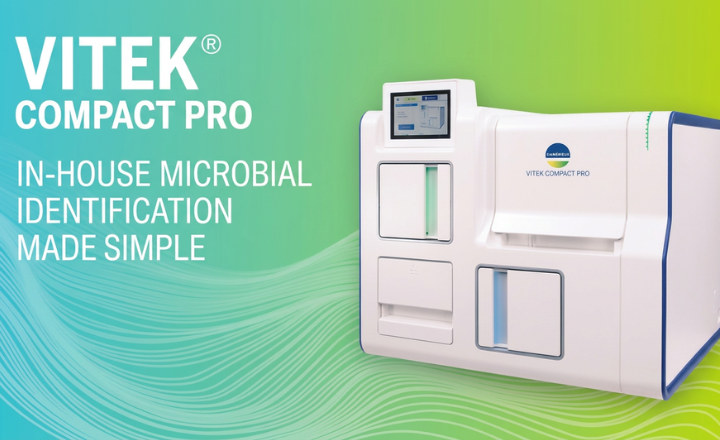
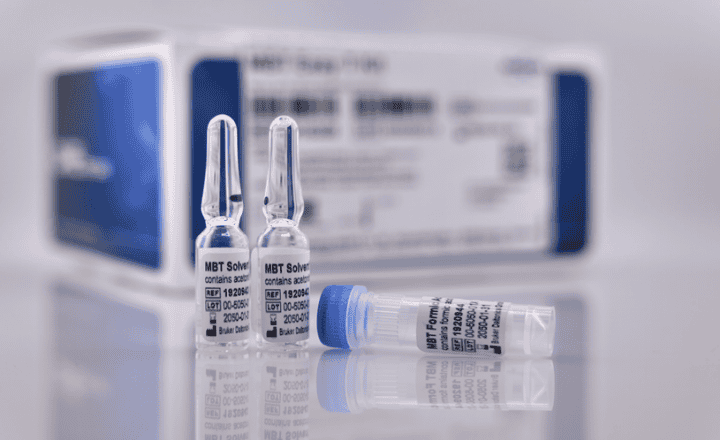
bioMérieux have launched the first CE-marked database and reagent kits for the identification of mycobacteria, Nocardia and moulds in a MALDI-TOF Mass Spectrometry System. This innovative solution further enhances the VITEK® MS system’s performance by adding 297 new species including identification of the Mycobacterium tuberculosis (TB) group, 45 species of the most frequent non-tuberculous mycobacteria (NTM), and 48 of the most medically important moulds. The VITEK® MS system’s new database and the reagents kits are now commercially available in countries which recognize CE-marking.
Mycobacteria, Nocardia and moulds are difficult to identify organisms, requiring days or weeks of specific culture conditions for appropriate growth and subsequent advanced methods for reliable identification to species level. The updated VITEK® MS system, however, can now offer simple, rapid and reliable identification of these pathogens, providing clinicians with actionable results to manage the infections which they cause, such as tuberculosis, lung and bone infections, and other serious organ infections.
Based on patented methods, the VITEK® MS Mycobacterium/Nocardia and Mould reagent kits enable easy, rapid, and safe identification of these organisms. The VITEK® MS extended database now enables the identification within minutes of 1,046 species representing 15,172 distinct strains of bacteria, yeasts and moulds. The convenient, user-friendly reagent kits for use with this highly automated mass spectrometry system provide lab professionals with extremely reliable results.
For microbiologists who choose mass spectrometry for microbial identification, bioMérieux offers a fully integrated solution combining identification with VITEK® MS and antibiotic susceptibility testing with VITEK® 2, resulting in enhanced pathogen information, and superior workflow management.