Creative Diagnostics...
bioMérieux La...
5th November 2020 Product update: rapidmicrobiology staff writer
Only Viable SARS-CoV-2 Material Detected With World-First Profiling Kit
Molecular diagnostics company Lucence has announced the availability of the world’s first assay kit to directly profile SARS-CoV-2 subgenomic RNA (sgRNA), a marker of active viral replication1, from clinical samples.
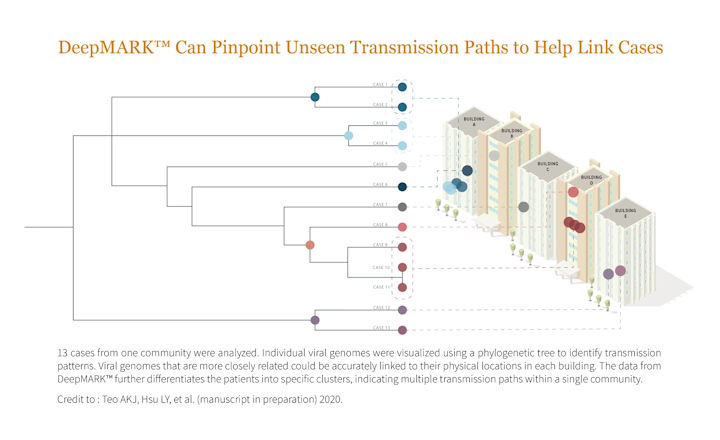
DeepMARK™ utilizes Lucence’s proprietary ultrasensitive next-generation sequencing (NGS) technology, AmpliMARK™, to concurrently detect and analyze the genome and transcriptome of SARS-CoV-2.
Pinpointing sources of unlinked SARS-CoV-2 cases supports rapid public health response. Using high-quality genetic fingerprinting, DeepMARK™ can enable rapid community case tracing by identifying transmission paths, clusters, and viral contagiousness.
DeepMARK™’s increased sensitivity also allows for asymptomatic and recovering cases to be more thoroughly evaluated for contagiousness.
For researchers studying contagiousness, SARS-CoV-2 sgRNA is a recognized marker of active viral replication linked with contagiousness2 and offers several advantages.
While viral culture is the gold standard, it is slow, expensive, and requires a biosafety level 3 (BSL3) laboratory, requirements prohibitive for the vast majority of samples.
Using DeepMARK™, clinical samples can be safely and efficiently profiled using a simple workflow.
SgRNA as a marker of viral replication is also a well-recognized efficacy measurement in SARS-CoV-2 vaccine development3.
Thus, comprehensive sgRNA profiling by DeepMARK™ could facilitate more efficient vaccine discovery.
“Genetic fingerprinting enables quicker and deeper analysis of viral spread. This tool has enhanced our capacity to understand SARS-CoV-2 precisely and contributes to the ongoing fight against COVID-19.” said A/Prof Hsu Li Yang, Vice Dean, Global Health, Saw Swee Hock School of Public Health, National University of Singapore.
DeepMARK™ is part of Lucence’s suite of SARS-CoV-2 molecular diagnostics. Lucence also makes the SAFER™ Sample Kit, a saliva stabilization kit with reported 36% higher sensitivity for detecting COVID-19 compared to nasopharyngeal swabbing4.
Tags:
Date Published: 5th November 2020
Source article link: View
Note: This content has been edited by a rapidmicrobiology staff writer for style and content.
Creative Diagnostics Supports COVID-19 Testing
bioMérieux Launches Predictive







