Article by Erin Patton, Senior Product Specialist – Charles River Microbial Solutions
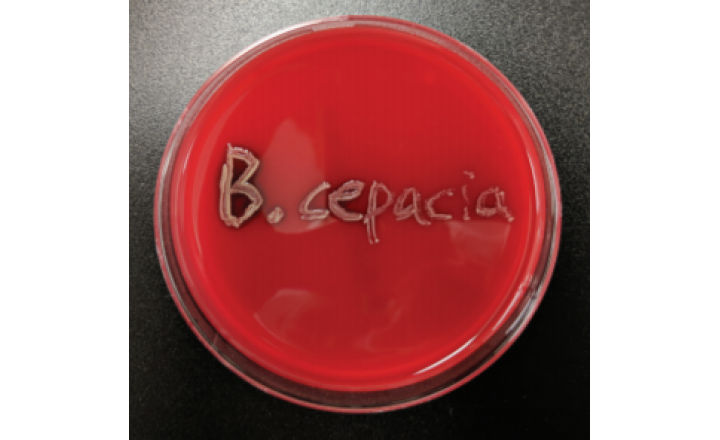
The Burkholderia cepacia complex (BCC) species are a group of gram-negative, rod-shaped bacteria which have been shown in recent years to be of concern for patients, and as such, for manufacturers of drugs and health care products that contribute to patient health. Species belonging to the BCC are opportunistic pathogens which have been involved in negative patient outcomes, especially for particularly vulnerable patient populations. As such, regulatory bodies tasked with protecting public health, such as the US FDA, have taken notice and issued numerous recalls for products that contain species from the Burkholderia cepacia complex. In turn, manufacturers have begun to revisit microbiological controls and their efficacy in preventing BCC-contaminated products. Aside from the need to detect the presence of B. cepacia complex species in tested samples, another important tool in the repertoire of microbiological controls is the ability to accurately identify microorganisms to the species level. B. cepacia species have proven problematic in this regard, challenging traditional methods and necessitating advances in microbial identification strategies.
It is critical that our academic knowledge about this complex of microorganisms continues to evolve as we also strive to evolve our knowledge of the clinical impact of BCC and the importance of microbiological controls against BCC in the manufacturing process. Clinically, BCC has been implicated for decades in outbreaks and cases of illness and even deaths1, 2. Members of the B. cepacia complex are often opportunistic pathogens among mechanically ventilated patients, the immunosuppressed, and those with serious underlying disease. Of particular note is the threat that BCC infections pose to patients with cystic fibrosis (CF). In a recent review, Sousa notes that BCC may cause serious infections in individuals with cystic fibrosis, and while all the BCC species may give rise to infection, the large majority of such infections are caused by two species in particular— Burkholderia cenocepacia and Burkholderia multivorans3. Further, Sousa notes that BCC infections are especially frightening for CF patients because of the antimicrobial resistance demonstrated by members of the B. cepacia complex, the highly contagious nature of BCC species, and the possibility that such an infection might give rise to additional complications like cepacia syndrome. Cepacia syndrome is developed in 20% of cases and is characterized by a swift, and often deadly, necrotizing pneumonia along with septicemia, sometimes progressing to the point of death in a matter of mere days.
Though B. cepacia is certainly of great concern for those who are immunocompromised or living with cystic fibrosis or chronic granulomatous disease, there are occasional cases in which B. cepacia has caused serious illness in non-immunocompromised, previously healthy patients4. BCC is highly contagious and especially virulent. With this risk to patients in mind, in recent years regulatory bodies like the FDA have taken a close look at products which contain BCC, issuing recall after recall and even an advisory statement to drug manufacturers in May of 20175, warning of the particular risk posed by BCC in non-sterile water-based products.
Non-sterile water-based products have featured prominently in BCC-related recalls, including products ranging from sanitizers, oral pharmaceuticals, gas relief drops, eyewashes, nasal sprays, mouthwashes, anti-cavity rinses, skin creams, baby and adult washcloths, surgical prep cloths, electrolyte solutions, and radiopaque preparations. In products such as these, BCC is especially troublesome because of its ability to resist preservatives and antimicrobial agents, growing even in unfavorable environments. One species in particular, B. multivorans, has been noted to grow well in low-nutrient environments such as distilled water, and is a good biofilm former. A recent case study shared by the FDA examined a situation in which B. multivorans was detected in a product (albeit prior to release of the product by the manufacturer). As it happened, two batches of a nasal spray containing an antimicrobial preservative were discovered during microbial testing to contain Burkholderia cepacia complex. Retesting other batches revealed that five additional lots contained BCC as well, though they previously had tested negative. The firm established that bacterial growth was initially inhibited by preservative in the product, but later overcame the preservative system and flourished. Detailed testing to elucidate the exact BCC species responsible for the contamination revealed Burkholderia multivorans as the culprit. Identification of the microorganism not just as a member of the B. cepacia complex, but specifically as B. multivorans allowed the firm to link the batch contaminations to each other and to perform a high-quality internal investigation to find the source of the contamination. Ultimately, the organism was traced back to a B. multivorans biofilm which had formed in the purified water system, and having established root cause, the firm was able to correct the plumbing and continue with additional corrective and preventative actions (CAPAs).
Some might suppose that it ought not matter to define exactly which BCC species is in question in such a situation, but in fact, it is important to know which particular species is implicated. It is not enough to assume that any one measure (CAPA) is sufficient to eradicate all different species, as they have distinctive characteristics and may respond differently to attempts to decontaminate. Additionally, as mentioned earlier, some species like B. multivorans demonstrate exceptional virulence, and may merit extra concern, action, or a broadened scope of investigation should they be uncovered in products, especially those which might be encountered by susceptible populations. Important though it is for the industry to be able to define the different species that make up the B. cepacia complex, it’s a difficult prospect to do so. At present, speciation of BCC is problematic, with many available methods yielding incorrect results or being incapable of distinguishing one species from another within the complex altogether6.
With respect to more advanced methods such as genetic sequencing, reliance on typical 16S sequence data is not a fully viable option either, as the species of BCC are so closely related that they are not distinguishable from each other in these genetic regions. Historically, the first 500 base pairs of the 16S rRNA gene have been used for sequence-based identifications of microbial samples sourced from manufacturing facilities’ environmental monitoring programs. This approach has shown to be very successful in providing identification results to the species level. But, even 16S rRNA gene sequence analysis has limitations with differentiating some groups of closely related organisms that share a high percentage of 16S rRNA gene sequence similarity, B. cepacia complex included.
As such, the pharmaceutical industry has taken a recent interest in the possible use of the entire 1500 base pairs of the 16S rRNA gene versus just the first 500 base pairs as is current common practice in bacterial identification. To evaluate whether this full-length gene sequencing might be a more appropriate approach, especially in situations wherein species cannot be differentiated with only 500 base pairs of sequence, a study was carried out by Hong and Farrance7. Findings showed that the overall performance of the first 500 base pairs of sequence of the 16S rRNA gene was quite high. When compared to the full-length 1500 base pair sequence for identification, 93.7% of the samples showed no differences between the two approaches. Considering the comparable performance of both 500 base pair and 1500 base pair sequencing identifications, as well as logistical limitations of generating the full-length sequence data for the 16S rRNA gene, it was concluded that in cases in which additional resolution is needed for confident speciation, it would make more sense scientifically and operationally to target alternate regions of the genome.
One proposed alternate gene target for providing the genetic discrimination needed to reliably differentiate closely-related species, including BCC species, is a protein-coding gene called recA. recA codes for a protein essential for repair and recombination of DNA8, and its sequence data have proven very valuable for the identification of B. cepacia complex species9 as well as other closely-related bacteria10. recA is uniquely suited to discernment of related BCC species as it exhibits higher sequence heterogeneity than 16S rRNA, yet is still highly conserved. Typically, members of the same BCC complex species demonstrate 98-99% sequence similarity at the recA target, whereas sequence similarity between different species drops to 94-95%11.
Recognizing the advantages presented by the recA sequencing approach when attempting to speciate members of the B. cepacia complex, service laboratories such as Charles River have developed the capability of distinguishing BCC species from one another with services like Charles River’s ProSeq offering. Because of the novel and complex approach in targeting recA, it is imperative that service providers use high quality sequence data, curated reference databases and updated bacterial nomenclature and taxonomy for phylogenetic analyses in order for an identification to be accurate and reliable. Charles River Labs puts all these safeguards in place as they recognize that accurate and reliable microbial identification data is critical not only for certain organisms such as the Burkholderia cepacia complex, but for all microbial isolates that are part of quality control practices.
References:
- www.cdc.gov/hai/outbreaks/b-cepacia-saline-flush/index.html
- www.ncbi.nlm.nih.gov/pmc/articles/PMC1490838/
- Sousa, SA et al., Burkholderia cepacia complex regulation of virulence gene expression: A review; Genes, Vol. 8, No. 43 – 2017
- Hobson R, Gould I, Govan J. Burkholderia (Pseudomonas) cepacia as a cause of brain abscesses secondary to chronic suppurative otitis externa.Eur J Clin Microbiol Infect Dis 1995;41:908–911.
- "FDA Advises Drug Manufacturers that Burkholderia cepacia complex poses a contamination risk in non-sterile, water-based drug products” (https://www.fda.gov/Drugs/DrugSafety/ucm559508.htm)
- Mahenthiralingam E. et al. The multifarious, multireplicon Burkholderia cepacia complex. Nat Rev Microbiol. 2005 Feb;3(2):144-56.
- Hong and Farrance. www.americanpharmaceuticalreview.com/Featured-Articles/181783-Is-it-Essential-to-Sequence-the-Entire-16S-rRNA-Gene-for-Bacterial-Identification
- Mahenthiralingam E. et al. DNA-Based diagnostic approaches for identification of Burkholderia cepacia complex, Burkholderia vietnamiensis, Burkholderia multivorans, Burkholderia stabilis, and Burkholderia cepacia genomovars I and III. J Clin Microbiol. 2000 Sep;38(9):3165-73.
- Pimentel, Dubedat, Dodds, Benn. J. Clin. Microbiol. November 2007 vol. 45 no. 11 3853-3854.
- Blackwood K. et al. Evaluation of recA Sequences for Identification of Mycobacterium Species. J Clin Microbiol. 2000 Aug; 38(8): 2846–2852.
- Vandamme, P and Dawyndt, P 'Classification and identification of the Burkholderia cepacia complex: Past, present and future' Systematic and Applied Microbiology 34(2)(2011):87-95
 About the author: Erin received her Bachelor of Science degree in biology from Xavier University and her Master of Science degree in cell and molecular biology from the University of Illinois, Urbana-Champaign. Erin has over 14 years’ experience in molecular biology research, microbiology QC, and regulatory compliance. As a Senior Product Specialist for Charles River’s Microbial Solutions division, Erin provides innovative rapid micro solutions for endotoxin testing, microbial identification, and microbial detection She is based in St. Louis, Missouri.
About the author: Erin received her Bachelor of Science degree in biology from Xavier University and her Master of Science degree in cell and molecular biology from the University of Illinois, Urbana-Champaign. Erin has over 14 years’ experience in molecular biology research, microbiology QC, and regulatory compliance. As a Senior Product Specialist for Charles River’s Microbial Solutions division, Erin provides innovative rapid micro solutions for endotoxin testing, microbial identification, and microbial detection She is based in St. Louis, Missouri.