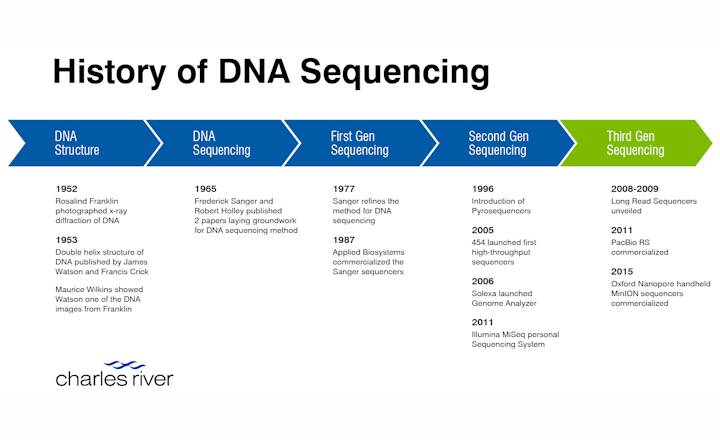
Tracing the history of DNA sequencing from foundational discoveries to current rapid advancements.
The Foundations of DNA Sequencing: 1871-1952
The history of DNA sequencing can be traced back to the late 19th century when scientists like Friedrich Miescher, Walther Flemming, Albrecht Kossel, and Phoebus Levene laid the groundwork in cell chemistry and nucleic substances. By 1929, the chemical composition of nucleic acids and the structure of DNA's backbone were understood, revealing the existence of nitrogen-containing bases (A, C, G, T, and U) and the alternating sugar-phosphate backbone.
The Revolutionary Discovery: 1944 - 1952
In 1944, Oswald Avery, Colin MacLeod, and Maclyn McCarty's groundbreaking research demonstrated that DNA, not protein, carries genetic information and can transform cellular properties. This discovery led to a significant shift in scientific focus towards DNA research. In 1952, Erwin Chargaff and Linus Pauling made important contributions by revealing crucial chemical properties of DNA, including the base-pairing rules (A=T and C=G).
The Sanger Method: 1965 - 1982
The first sequencing methods were introduced in the 1960’s, with a heavy focus on RNA sequencing due to its relative simplicity. It was not until 1977 that Frederick Sanger developed a method for DNA sequencing that became the gold standard for over a decade. Sanger's method enabled a major milestone, the sequencing of the first whole genome of a bacteriophage in 1982.
The Human Genome Project: 1984 - 2003
In 1984, the idea of sequencing the human genome to identify disease-associated genes was proposed. In 1990, the Human Genome Project was launched, aiming to sequence the entire human genome. After 13 years of collective effort from the international Human Genome Sequence Consortium, the first draft of the human genome was published in 2001, and the final version was released in 2003.
Second-Generation Sequencing: 1996 - 2006
The emergence of second-generation sequencing technologies began with the introduction of pyrosequencing in 1996. These methods, including Roche/454, SOLiD, and Illumina, allowed for high-throughput sequencing and significantly reduced the associated costs. Roche/454 Life Sciences released the first high-throughput sequencer in 2005, followed by Illumina's Genome Analyzer in 2006, propelling the field of genomics into a new era.
Third- and Fourth-Generation Sequencing: 2008 - Present
Third- and fourth-generation sequencing technologies revolutionized DNA sequencing by providing long read capabilities. Pacific Biosciences's SMRT sequencing uses fluorescent nucleotides to generate long reads, and Oxford Nanopore Technologies 's nanopore technology measures disruptions in electrical current as individual bases pass through a nanopore. These technologies have facilitated de novo genome assembly and structural analysis, allowing researchers to study previously unexplored organisms and improve disease screening methods.
Current Advancements in Microbial Identification
Advancements in sequencing technologies have also transformed microbial identification. Previously, DNA-DNA hybridization was used, but next-generation sequencing techniques, like whole genome sequencing and ONT sequencing, have enabled more accurate and strain-level microbial identification.
The history of DNA sequencing is a testament to human ingenuity and dedication to unraveling the mysteries of life. From 13 years to 6 hours…the speed and efficiency of DNA sequencing has come a long way. Learn more about the fascinating advancements in sequencing technology and methodologies in this blog post.
Read the full article here or use the Request Information button to send an email.