Karin Pawlowsky, Microbiology Manager, Campden BRI shares with rapidmicrobiology.com her knowledge of rapid test methods for brewing microbiology.
Beer is relatively microbiologically stable. This is due to five inherent properties of beer:
1. The alcohol content acts as an antimicrobial 2. Iso-α-acids, which are derived from hops, also inhibit microbial growth 3. Low pH 4. The nutrients, that could support microbial growth, are depleted after fermentation 5. The oxygen content of beer packaged in bottle, can or keg is very low which limits the growth of aerobic organisms.
In combination, these five microbiological hurdles mean that microbial survival/growth in beer is limited and only those microorganisms able to overcome the hurdles can pose a risk to product quality. Pathogenic organisms generally do not survive in beer so brewers are mostly concerned with spoilage organisms of which the most problematic are anaerobic bacteria and fermentative yeast species.
Brewers don’t usually just check their final beer for spoilage microorganisms, but also monitor brewing samples to ensure that the process is contamination free. Process samples may include wort, pitching yeast as well as fermentation/maturation/bright beer tank samples.
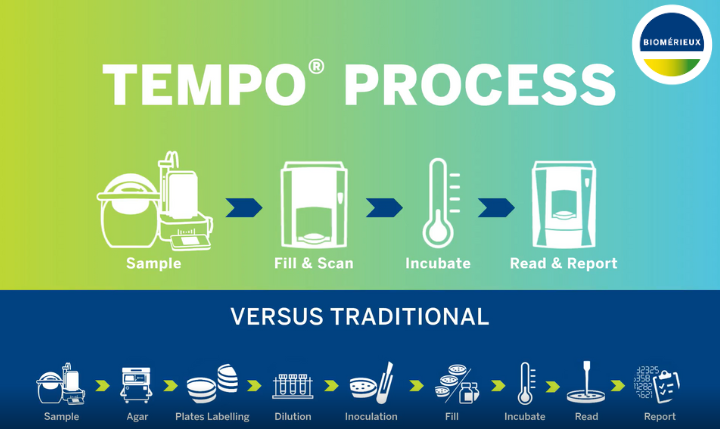
Traditional Microbiology for Brewing Brewery samples are often screened for spoilage microorganisms using traditional plate-based microbiological techniques. A number of growth media are used for this purpose, some of which have been specifically developed for the brewing sector. Typically, samples are screened for aerobic yeast and bacteria as well as anaerobic bacteria. Membrane filtration is the method of choice for filterable samples resulting in test sensitivity of 1 cfu per 100ml. Non-filterable samples are commonly spread plated reducing the limit of detection to 10 cfu/ml.
Anaerobic plates are usually incubated for 5-7 days whereas the aerobic plates only take 3-5 days for colony formation. The pattern of growth on the various media, some of which are selective or differential, together with the colony morphology give a good indication of the type of microorganisms present. In many cases this information is sufficient for brewers and further identification may not be required. For example, should anaerobic growth be observed this is most likely to be due to lactic acid bacteria which are considered potential beer spoilers and would require action to be taken.
However, in some cases further analysis of the growth on plate can be helpful. A portion of a colony is examined microscopically to determine the organism’s cell and group morphology. Additionally, Gram staining, catalase and oxidase testing can be performed if necessary. In combination, these results give a good understanding of the family or genus of any microorganisms present and at which concentration. Should there be a requirement for an exact identification to genus level further analysis would be required. Although traditional brewing microbiology yields good results the major disadvantage is the long plate incubation period. Additionally, some organisms cannot be detected such as strict anaerobic beer spoilers (Pectinatus spp., Megasphaera spp.).
Rapid Microbiology for Brewing New rapid technologies have been developed to enable brewers to act more quickly on microbiological issues. Many of these are DNA-based and employ PCR to amplify a specific DNA target sequence which then allows detection of the target. A number of commercial PCR test kits designed specifically for brewing applications are available from various suppliers. These kits can screen samples to cover a group of organisms (e.g. lactic acid bacteria) or identify to species level. The kits can provide results within a few hours if sufficient microbial cells are present. If this is not the case, samples need to be pre-enriched, which adds~24-48 hrs to the testing time. A disadvantage of this technology is that there is no differentiation between live and dead cells so false positive results are possible.
Furthermore, enumeration is not feasible; it is very much a presence/absence test. However, due to the speed of the tests and ability to detect the strict anaerobic beer spoilers, these rapid technologies are being employed by some brewers. In the last few years there have been developments to simplify result interpretation (e.g. lateral flow assays) and minimise sample handling (e.g. fully automated system;). Some of these later versions also include testing for hop-resistance genes so beer spoilers can be differentiated from non-beer spoilers.
Rapid detection of organisms present in beer can also be done using other techniques, such as micro-colony counting. Growing microbial colonies are stained, then detected and counted before they are visible to the eye. Slow-growing lactic acid bacteria can be enumerated within three days. Alternatively, live cells can be detected by methods employing in situ hybridisation, the binding of labelled DNA probes to a target RNA sequence. Cells are bound to a slide, their cell membrane enzymatically degraded, then treated with fluorescent labelled probes and incubated (hybridisation). The sample is finally examined for live cells by fluorescence microscopy. Another variation of this technology uses sandwich hybridisation where two probes are used: a capture probe that binds the target DNA to a microtiter plate well and a detection probe that visualises (via an ELISA type reaction) the bound target.
Matrix-Assisted Laser Desorption/Ionisation Time of Flight Mass Spectrometry (MALDI TOF MS) is now available for rapid identification of microbes. A laser is fired at the isolated cells which produce ionised proteinaceous molecules. These are accelerated through an electric field and hit a detector at different times according to their varying masses. This produces a species-specific fingerprint that can be compared to a fingerprint database of known species to identify the microbes.
More information about some of the technologies mentioned can be found in our compare products database:
About the author: Karin Pawlowsky has over 10 years of experience of microbiology. Karin’s areas of expertise are: brewing microbiology, turbidity in beverages and dispense hygiene. Her team works with member and non-member companies worldwide on troubleshooting and project work, including grant funded projects. In her time at Campden BRI Karin has presented at many events and has also delivered numerous training courses.