by Vikki Mitchell, Identification Services Manager, NCIMB
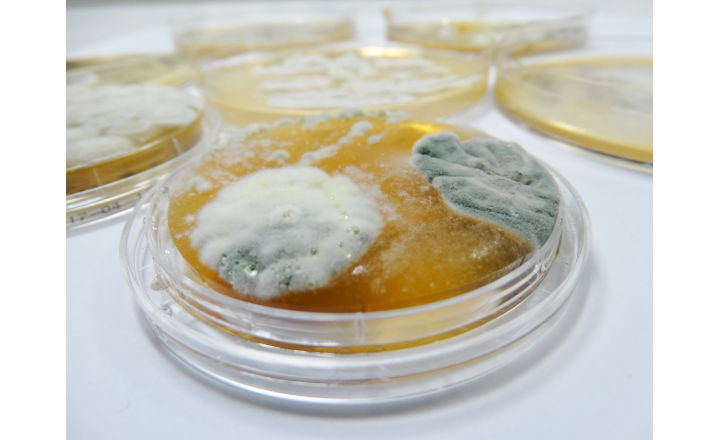
Species level identification of fungi has long been considered challenging and consequently, in contrast to bacteria, fungal isolates obtained from environmental monitoring programmes are often identified to genus rather than the species level. More detailed identification of environmental isolates can be desirable - so what are the difficulties when it comes to species level identification of fungi?
Bacteria are commonly identified by sequencing of the 16S ribosomal DNA, and the approaches that have been developed for genotypic identification of yeasts and moulds also involve sequencing sections of ribosomal DNA. Fungal ribosomes have a large and small subunit. The ribosomal RNA operon, that is the DNA that codes for ribosomal RNA, has three rRNA sequences and two internal transcribed spacer regions: ITS1 and ITS2.
Two distinct approaches have emerged for genetic identification of fungi: sequencing of the D2 region of the large subunit ribosomal DNA (D2 LSU) and sequencing of either one or both of the internal transcribed spacer regions (ITS).
D2 LSU sequencing is probably the most widely used approach for fungal identification at present. Similarly to 16s rDNA sequencing for bacteria, data obtained can be analysed using a validated commercial database that has been built using reference strains, as well as publicly available but non validated sources such as the European Nucleotide Archive (ENA) database from EMBL-EBI (European Molecular Biology Laboratory - European Bioinformatics Institute). The ITS approach however, generally relies more heavily on the use of data obtained from unvalidated public databases.
While I find it is usually possible to obtain a species level identification of yeasts using D2 LSU sequencing, we sometimes can’t get that level of identification for moulds and in these cases, it is often possible to obtain species level identification using ITS sequencing. Generally, there is a higher level of differentiation between ITS sequences. Unlike ribosomal DNA, ITS sequences have no functional role, and consequently have accumulated a greater level of mutation, which aids identification.
A good example of moulds that are difficult to identify to species level using D2LSU comes from the genus Penicillium - a commonly occurring genus amongst environmental isolates. I have found Penicillium camemberti, Penicillium clavigerum, Penicillium commune, Penicillium corylophilum and Penicillium crustosum have all matched a single isolate sequence at 100% similarity. However, when ITS sequencing has subsequently been undertaken, a species level result has been achieved – in one specific example the isolate was identified as Penicillium crustosum with a 100 % match
The above example is an illustration of where D2LSU sequencing cannot differentiate between different species of the same genera, but in some cases, D2LSU sequencing alone cannot distinguish between different, but closely related, genera. For example, I have found several species of Cladosporium and Mycosphaerella have all matched to the same isolate sequence. Cladosporium is a large genus that has been reported to be the most common fungal component isolated from air, and is therefore also quite commonly found in the course of environmental monitoring programs. Again, at NCIMB we have found ITS sequencing to be successful in providing a species level identification - one isolate which matched to Cladosporium cladosporioides, Cladosporium herbarum, Cladosporium oxysporum, Mycosphaerella aronici and Mycosphaerella tassiana, was identified as Cladosporium cladosporioides when ITS was used.
In another example, we found that it was not possible to differentiate between an even larger group of closely related genera. Searching the non-validated EMBL database for D2LSU sequences failed to distinguish between Saccothecium sepincola, Mycosphaerella sojae, Pithomyces chartarum, Pleospora gaeumannii, Leptosphaerulina saccharicola, Heterophoma adonidis, Nothophoma quercina and Leptosphaerulina australis . These eight species, from seven different genera, all matched a single D2LSU isolate sequence at 100% similarity. In this case however, ITS sequencing did not result in a species level match either – the isolate matched to Leptosphaerulina saccharicola, Leptosphaerulina australis, Leptospherulina chartarum and Leptosphaerulina trifolii . It was, however, successful in narrowing the identification down to Leptosphaerulina species rather than several different genera – a substantial improvement on the initial result obtained.
As many fungi do match very well to the validated D2 LSU database, at present, we recommend the use of D2 LSU sequencing in the first instance. If a match is not found using the validated database, we would analyse the results against the non-validated ENA database, before considering whether to follow up with ITS sequencing. We always refer to any relevant published papers for additional supporting information when using unvalidated databases for identification purposes.
The decision on whether to progress to ITS sequencing if a species or genus level identification cannot be obtained using D2 LSU sequencing is really dependant on the individual circumstances and whether family, genus or species level identification is required. For example, it may be requested when investigating excursions from normal populations or contamination issues. While ITS does not always provide a species level match, the examples above illustrate that it has been successful in doing so with some commonly isolated fungi, and where a species level match has not been found it has given an improved result. Validated databases of ITS sequences are being developed and in future, our experience suggests that this may result in increased popularity of the method.
Vikki Mitchell is available to answer your questions on species level identification of fungi. Use the 'Request More Information’ button below.
About the author: Vikki Mitchell joined NCIMB in 2005. She leads a team of scientists responsible delivering NCIMB’s Identification Services and sequencing new deposits to the UK’s National Collection of Industrial Food and Marine Bacteria. Vikki holds a BSc (hons) degree in Applied Biosciences and Management, and an MSC in Instrumental Analytical Techniques; DNA Analysis, Proteomics and Metabolomics from the Robert Gordon University in Aberdeen.