Why This Matters:
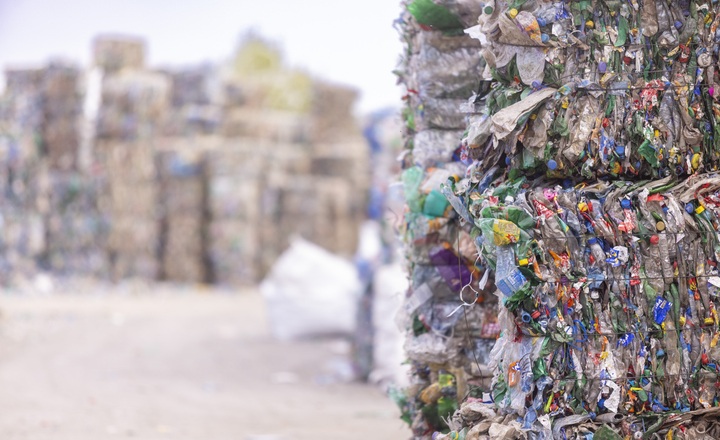
- Occupational health risk: Waste sorting workers are exposed to complex bioaerosols that include microbes and microbial fragments, which may contribute to respiratory disorders, allergic reactions, or infections.
- Limitations of traditional methods: Culture-based microbiology misses the large fraction of non-culturable organisms, underestimating biodiversity and potential hazards.
- DNA metabarcoding power: HTS of amplicon markers (16S rRNA for bacteria and ITS for fungi) offers species-level resolution, facilitating detection of human pathogens and multi-drug resistant taxa within bioaerosols.
- Evidence for risk profiling: Identification of genera such as Aspergillus (including A. fumigatus), Fusarium, Candida, Enterobacteriaceae, and Staphylococcus aureus suggests potential occupational exposure to clinically relevant microbes.
Key Findings: Eriksen et al. used high-throughput sequencing (HTS) to characterize the airborne microbiome across six waste sorting plants, aiming to better define the biological exposures faced by workers in these types of facilities.¹
- Microbial community composition: Personal air samples contained >2,000 unique bacterial and fungal amplicon sequence variants (ASVs), with bacteria dominating (~68%) followed by fungi (~32%).
- Diversity variation: Alpha diversity (richness and evenness) varied significantly between plants and with certain work tasks that generated more dust, suggesting task-dependent exposure profiles.
- Dominant taxa: Cladosporium species dominated the fungal fraction, while Aerococcus and diverse bacterial genera were prevalent; community composition differed substantially between facilities.
- Presence of pathogens: Sequencing detected species associated with human disease — e.g., Aspergillus fumigatus, Candida albicans, Fusarium spp., Staphylococcus aureus, and Enterobacteriaceae.
- Seasonality and environment: No consistent seasonal effects on diversity were observed; variability was more closely linked to waste material and operational differences rather than temperature.
Bigger Picture: Integrating DNA metabarcoding into occupational hygiene frameworks can substantially improve microbial risk assessment by capturing both culturable and non-culturable organisms with species-level precision. This facilitates identification of hazardous taxa that conventional methods overlook, supports targeted exposure mitigation (e.g., ventilation or protective equipment during high-dust tasks), and can inform regulatory guidelines for worker safety. Continued validation across broader geographic settings, sequencing depths, and longitudinal monitoring will be needed to better translate these molecular insights into health risk thresholds and intervention strategies.
(Image Credit: iStock/ Lajst)