Recently returned from the Marijuana Business Conference and Cannabis Expo in Las Vegas, Milan Patel CEO of PathogenDx, talks to rapidmicrobiology.com on why the ultra-fast PathogenDx molecular-based pathogen testing platform is being adopted by state-licensed cannabis testing laboratories in the absence of federal regulations and how disruptive the technology could be to gold-standard plate testing in food, water and environmental testing laboratories.
Q: What is the technology behind PathogenDx?
A: PathogenDx is both a qualitative and quantitative test for not only pathogenic species but also class indicator organisms. The basis of it is an enrichment-free DNA molecular platform that can test for multiple organisms with the aid of a microarray. A microarray is essentially a microscopic glass slide with wells that contain the arrays. The arrays are essentially probes that are printed that characterize the DNA sequence of up to four dozen pathogens, both bacterial and fungal in triplicate. Each probe represents one of the DNA complementary pair for an individual organism or analyte. A fluorescent LED imager identifies and quantifies the pathogen profile when the arrays are imaged, and with the aid of cloud-based software, results are delivered within 6 hours.
Q: What are the steps involved in identifying and quantifying pathogens using PathogenDx?
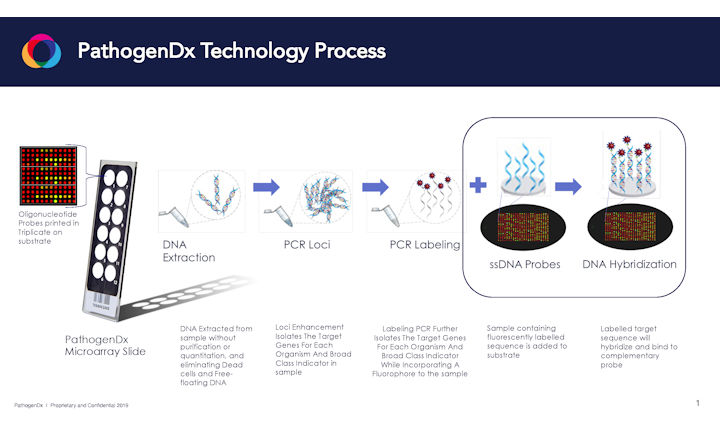
A: Whether its food, agricultural or botanical material you're testing, the same procedure applies: You effectively take the matrix you are testing, and take it through a very simple sample prep, lysis process. Then through a tandem PCR process you drop the post-PCR processed sample onto the microarray, letting it hybridize, washing it and then imaging the slide and letting the software do the rest to get the results.
The sample prep and lysis process calls for centrifuging the matrix material into a very small pellet: and using this as your sample input that contains the bacterial, fungal or viral cells that will undergo the rest of the testing steps.
In terms of sample prep, the matrix material is placed in PBS, and then it is vortexed. After that, some of the fluid is drawn and sent through a two-step centrifugation process. The first step requires 50 x g centrifugation to send the dense matrix material to the bottom, leaving a supernatant to pipette out. For the 2nd step, in order to get to the pathogenic cells, the supernatant is spun at a much faster 14000 x g, which sends the higher density live cells to the bottom in the form of a 50µl pellet, leaving the dead cells on top.
Then the pellet is sent through a sample lysis process with a proprietary digestion buffer, which heats up and cracks open the pathogenic cells, allowing you access to the DNA. This DNA is then used in a 1- hour enrichment-PCR step, and then a 1-hour labelling-PCR step, which involves attaching a fluorophore to the pathogen or analyte DNA.
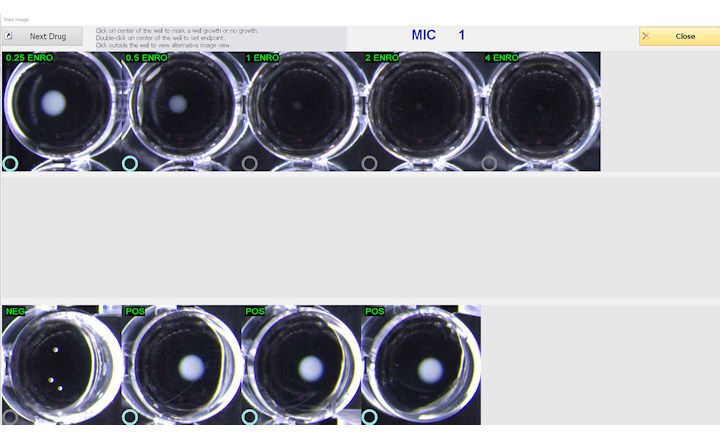
The sample is then placed onto one of the microarray’s wells, and allowed to sit for 10 minutes, to allow hybridization of the sample’s DNA, binding to its complementary pairs in the well. This microarray is then placed in a fluorescent LED imager, which excites the probes, and the image produced is sent to the cloud with the software, generating an identification and quantification report based on the microbial load of the pathogens in that sample.
You can do a full 96-well plate or 96 samples within 6 hours with only an hour and a half of hands-on time.
Q: How is PathogenDx able to detect down to 0.1 CFU?
A: We have been able to put super high-sensitive probes onto the microarray that work well with the primers we designed. That particular primer-probe combination allows for a superior sensitivity as well as a specificity. To achieve this, we performed a logarithmic dilution using purified DNA from 50,000 femtograms (10-15) down to 5 femtograms, where one CFU represents between 5-50 femtograms of the genomic DNA. So as we were able to get detection below five femtograms that gives us one log below that one CFU level. This is all based upon the most probable number technique to a large degree.
Q: What range of organisms can the PathogenDx detect?
A: It can detect bacterial, fungal and viral, and because it's DNA based, you can identify stressed organisms and also viable and non-culturable as well (VBNC), unlike plate culture. We can detect and quantify fungi by using the ITS1, and ITS2 region of the genome and bacteria are identified by the 16S section of the genome. We also plan to have a plant virus multiplex assay that will be introduced in the first half of 2020 to tackle the range of viruses, like the tobacco mosaic virus (TMV) that have been a huge issue in the agricultural sector.
Q: How does this compare to other PCR kits on the market?
A: The first discriminator is that other PCR kits require sample enrichment, DNA extraction, purification and quantitation steps along with a sample preparation kit. Pathogen Dx has developed a technology called ‘raw sample genotyping’ (RSG) which obviates the need for DNA extraction, purification, quantitation and the enrichment step. The second major discriminator is that we can identify or detect and quantify microbes in a multiplex.
So what does that mean? That means, in other words, all of the organisms you are looking for in a sample are in the same reaction. And so other PCR kits must use a single plex approach, for instance, they would have to test for E. coli and Salmonella separately, requiring individual primers, along with the positive and negative controls, which increases the number of steps, complexity and cost. With our multiplex assay, you do an all-in-one reaction by using one universal primer and the controls built into the actual array. This reduces the number of steps a lab technician must execute; it brings down the cost and also reduces the turnaround time. You can run near to 100 samples, with each sample been tested for up to 50 pathogens in parallel.
Q: Why has PathogenDx not already been introduced into the food industry?
A: One of the challenges we faced was the technology’s disruptive and innovative nature and how it would have been difficult to get the regulatory bodies to adopt such technology. So instead we went “under the hurdle” and entered the cannabis market, to show to the world the platform delivers results in 6 hours with superior specificity and sensitivity and at a lower cost. So now we have close to 100 customers using the platform day in, day out, as their primary microbial testing platform. There are no federal regulations, only state agencies, and that’s what was the input to our genesis; it would be a lot easier to get through the state-level regulatory bodies of which we’ve done, and so far we have the technology being used in about 20 different states. I think once cannabis gets de-scheduled and the FDA looks at our platform, I’m sure the process for federal-level regulatory approval in markets such as food will be easier given that it has been in use and used both nationally and internationally, its adoption in food will be easier.
Q: Who is allowed to do cannabis testing?
A: The states that are legalized either medically or adult-use or both, and have implemented a ‘seed to sale’ tracking system. They want growers to produce high quality and safe cannabis, and so they require a third-party independent certified laboratory, which is licensed through the state, to do the compliance testing of the product. Each sample goes from the grower straight to a lab, and the lab must then test, and submit COAs to the state as proof that a sample from a grower, processor or a producer complies to the state regulations for microbial content before that product can be released to a dispensary for consumption to a consumer or a patient.
Q: Do you think PathogenDx could have prevented outbreaks like E. coli 0157 in Romaine Lettuce?
A: Absolutely! When you look at the food or agricultural industry; products are moving from farm to fork at an accelerated pace. These industries have taken to using plate culture, which is the gold standard in classical microbial testing, but It’s a nineteenth-century technology. The challenges we have, you’ve seen it daily, whether its Romaine or chipotle 3 or 4 years ago, is that the velocity at which these industries and markets are operating at, with the food or product travelling through the supply chain is at a much faster rate. By the time produce like Romaine gets to a restaurant or a retail grocer, we’ve already consumed it, way before results from testing is available. And then its too late.
You need an ultra-rapid method with the superior resolution of specificity and sensitivity, which we have been able to provide with the microarray, to get that level of performance that’s similar to sequencing but not at the cost of sequencing but the cost of traditional microbial plate culture. We have delivered a perfect next-generation technology that fits within the detection limits that the food, agricultural, and water industries have set, so people don’t get sick or die. But can also give you an answer within one single lab testing shift. Whether you are at the farm or port of entry for food, or fruit and vegetable packing, you can run that test at that point and within six hours have the result and still be able to release the product knowing its absolutely safe and of the highest quality. That’s why I think this particular technology sits in a place of its own and can address some of the greatest challenges we are seeing, like almost hundreds of food-borne outbreaks a year just in the US alone.
Q: What regulatory approvals are in the pipeline for PathogenDx?
A: We are now entering the food world with AOAC approval of the technology expected in the first half of 2020. That’s going to be a proficiency test method, for both environmental and for specific food matrices. That approval will allow the technology to be used federally both in the United States and in the thousands of international food and environmental safety testing laboratories so that the technology can be adopted and used to address some of these challenges I’ve mentioned.
Q: If there was a PathogenDx 2.0, which parts of the technology might be improved?
A: The 2.0 would be a simpler version, reducing the turnaround time, even though its already only 6 hours, which is almost four times faster than anything out there. And also making it more point of application at the farm in a more easier handheld approach. There have been other companies that have tried to do that. Still, I think the biggest challenge in doing that is dealing with the matrix whether it’s a leafy produce matrix or meat, that in the sample prep itself we have to “crack that nut” and I think we have the team to address some of those areas. We’ve overcome the hurdle of ‘live vs dead’ cell detection testing with this platform; we’re doing multiplex pathogens, we’ve got superior sensitivity, the cost has been brought down to below or equal to plate culture, we do triplicate testing per analyte and so on. So we’ve knocked off some of the bigger technical challenges, and so the next version would be a mobile lab and having the capability to be handheld at some point.
Q: What is the future direction of PathogenDx?
A: Given the universal aspect of our technology, our tagline is 'to set the standard in DNA testing'. I believe the future is to move from the century of petri-dish testing into DNA based testing and to enter the food, agricultural and water industries and ultimately the healthcare markets. The protocol does not change whether you are in a food safety lab or an agriculture lab or a dairy lab or even a healthcare lab; it’s the same process. Let’s say we have ‘standardized’ the process no matter whether you are testing E.coli on cannabis, Romaine, feedstock, or human clinical sample. The only difference is what you start with as the sample, but every step after that remains the same. By doing this, it drives universal standardization – meaning the same process is administered– resulting in much greater efficiency, much better performance and lower costs than plate culture and other conventional DNA methods.
Q: Clostridium botulinum has been found on the cannabis plant, does PathogenDx test for the organism?
A: It's ironic you ask because the State of Florida had recently put Clostridium botulinum as a target for microbial testing halfway through 2019. Yes. The platform can test for this organism. We have the primers and the probes already on the detection array, and so you can test for it down to one CFU. For such a harmful organism, which even people say that you can eliminate during extraction, it's unknown at this point; you don’t want to take a risk, so we have the ability to test for it in our array.