The Oxford Nanopore handheld MinIOn sequencer has been evaluated for routine environmental monitoring at a food-processing facility.
Dairy processing environments harbor microorganisms that have the potential to contaminate food before and during processing. Some of these microorganisms have the potential to cause spoilage or be pathogenic. Many culture-based assays are unable to distinguish between undesirable taxa and closely related harmless species. Furthermore, even when multiple culture-based approaches are used in parallel, it is still not possible to comprehensively characterize the entire microbiology of a food-chain sample.
In addition, the high false-positive rate from some selective and phenotype-based agar assays was a given as a reason for doing the study.
Researchers from University College Cork (UCC) and Teagasc (Ireland's National Food Research Centre) took swab samples from 8 different locations at a dairy processing facility and used Nanopore's MiniION and Illumina NGS to provide microbiome data.
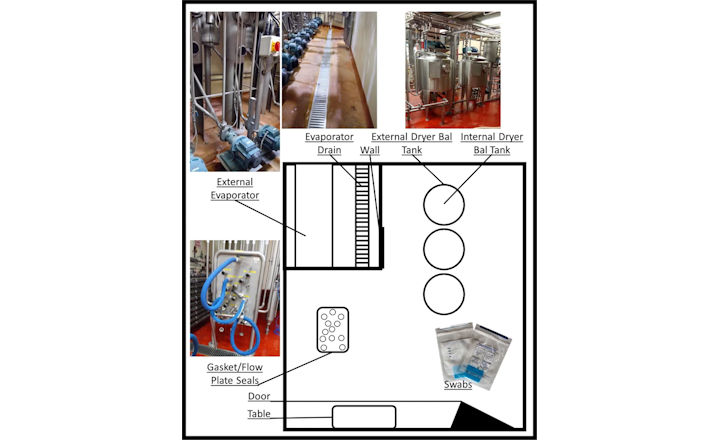
Swabs were taken from surfaces, which include: table, door, wall, evaporator, and dryer balance tank. Swabbing was carried out after clean-in-place (CIP) operations and before the next round of dairy processing.
High-throughput DNA sequencing represents a potential means through which microbial monitoring of the food chain can be enhanced. While sequencing platforms, such as the Illumina MiSeq, NextSeq, and NovaSeq, are most typically found in research or commercial sequencing laboratories, newer portable platforms, such as the Oxford Nanopore Technologies (ONT) MinION, offer the potential for rapid analysis of food chain microbiomes.
Overall, ONT MinION sequencing provided accurate classification to species level, which was comparable to Illumina-derived outputs. However, while the MinION-based approach provided a means of easy library preparations and portability, the high concentrations of DNA needed to run the rapid sequencing protocols was a limiting factor, requiring the random amplification of template DNA in order to generate sufficient material for analysis.
Click here to read study.
Note: This content has been edited by a rapidmicrobiology staff writer for style and content.