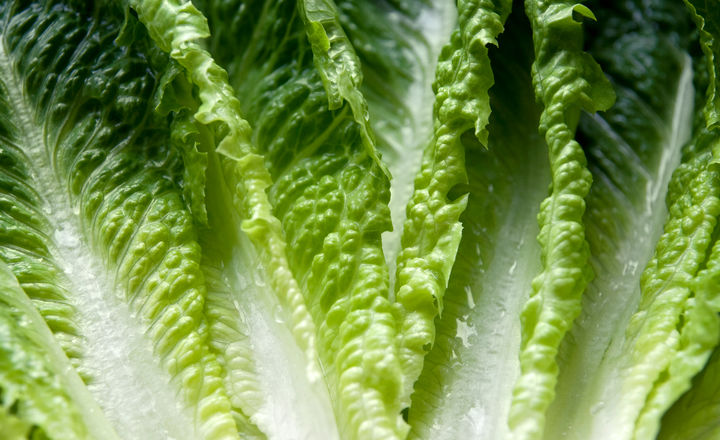
In April 2018, the CDC reported a multi-state outbreak of Escherichia coli O157:H7 infections linked to Romaine lettuce grown in Yuma, Arizona with five deaths so far reported and hundreds sickened. Here, rapidmicrobiology finds out from Dr. Rachel Noble, an expert environmental molecular microbiologist, how a recently developed molecular test for generic E. coli might be part of the solution.
How does E. coli O157 get onto leafy veg such as Romaine lettuce? E. coli are naturally part of the intestinal flora of cattle, poultry and other animals and can then make their way into the environment through manure present in fertilisers and leafy vegetables such as Romaine lettuce, grow right next to soil. There is also a possibility of O157 from biosolids; contaminated surfaces in packing and distribution houses or cross-contamination in restaurants serving the product.
Additionally, Romaine lettuce is heavily irrigated and in many states irrigation water is coming from quite a distance, in the form of pipes or even open canals so this is another potential source of contamination. Sometimes prior to the distribution of the lettuce, the produce is rinsed or even triple washed in a packing house. It is also possible that during this process contamination occurs from the water used. We, therefore, have various environmental sources that can contribute to heightened risk and as most people don’t cook their salads it results in the perfect ‘storm’.
To my knowledge, this outbreak involving Romaine lettuce has not yet been pinned to a single point in the process so the problem could be anywhere from out in the field to point of consumption. Looking back at recent outbreaks like the O157 outbreak from spinach in 2012, this ended up being traced back to run-off from a herd of cattle.
Are the current guidelines for detecting STEC sufficient? The detection systems do work but the system can be improved with an even more rapid response and potentially the use of more rapid molecular methods.
In the United States, there is a new set of rules associated with the FDA Food Safety Modernization Act, which includes the “Produce Safety Rule” which requires large farms to begin to test their agricultural water for E. coli throughout the year. I believe the implementation of this rule will help to identify problem areas and begin to collect data that identifies areas that might be high risk for contamination. This is still in the implementation phase and most farms have not fully begun to do their required testing and reporting.
As a researcher, I would love to see us take this proactive approach to determine the factors that promote microbial contamination in growing areas and distribution houses beyond not following basic hygiene standards. As data is generated and used to connect to the potential for outbreaks it won’t give us an opportunity to prevent all outbreaks, as the infectious dose for E. coli O157 is extremely low, but it will give us useful information that we currently lack.
Have the tracing and recall systems in place worked in this particular outbreak? In general, I can say the response has been rapid and well-coordinated across the agencies such as the FDA and CDC that we would want to be communicating. In the last ten years, I would say we have improved our tracking capability and shortened our response time. The fact that there is only one death from this outbreak with the rates of hospitalisations that occur for this organism, a rate between 30-50%, shows the media has been effective in alerting consumers to throw out the produce.
Tell us about the BioGX E. coli test you have been working on As my background is in environmental marine microbiology, initially the original intention was to produce more rapid test methods for recreational waters. I worked in collaboration with BioGX, who licensed a rapid test that quantifies all strains of E. coli. Then we realised there are other applications that we weren’t working on, one of them was analysis of produce water, the sample is essentially the same water with a little bit of sand and sometimes oil or sediment. This is how the produce application came about. We now do work on shellfish such as oysters and clams and so forth.
The test is presented in different formats, but it’s based on a real-time PCR approach with an easy to set up and run style. This means E. coli detection times are reduced from 24 hours+ to one to two hours. We are now actively working towards validation of this test for the generic E. coli testing requirements that are part of the Produce Safety Rule. A format of these tests is currently in use by various produce packers on the West Coast and it takes less than two hours to get results making it ideal for the rapid response times needed for this short shelf-life produce product.
This test can also be paired with other more specific tests for other pathogens such as Salmonella, to provide a rapid result on the presence of both in a produce wash water sample for example.
Dr. Rachel Noble is a renowned environmental microbiologist from the University of North Carolina at Chapel Hill. A main thread of Dr. Noble’s work is the application of novel molecular techniques for applied and basic science. She is currently working with BioGX developing a molecular kit that will test for three bacteria (E. coli, Listeria, and Salmonella) simultaneously. While Dr. Noble’s previous research studies have centred around developing rapid testing kits for water and seafood safety, her work has also been found to be applicable for testing agricultural produce as well, especially leafy greens like lettuce and spinach.