Article by Dr. Tim Sandle, Head of Microbiology at Bio Products Laboratory, UK
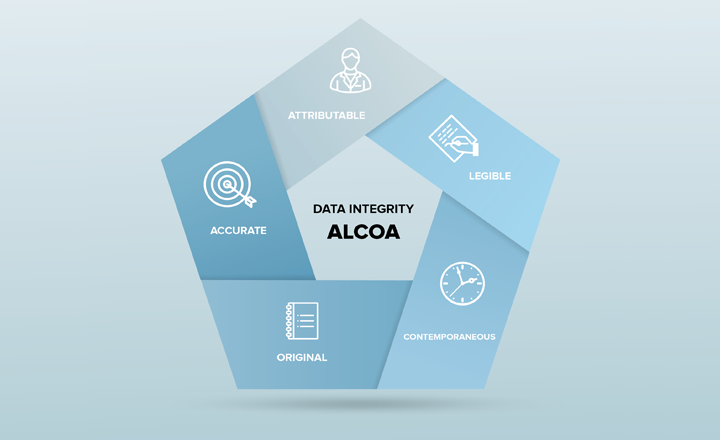
Data integrity refers to maintaining and assuring the accuracy and consistency of data over its entire life-cycle, and is a critical aspect to the design, implementation and usage of any system which stores, processes, or retrieves data. Data integrity is a key regulatory concern and to aid those working in pharmaceuticals and healthcare guidance documents have been produced by the U.S. Food and Drug Administration (FDA) and the MHRA (1, 2). These call on the ALCOA approach to be followed: data must be attributable, legible (permanent), contemporaneous, original and accurate.
Data integrity applies to all production processes and laboratories, including microbiology. That microbiological practices are a potential concern is evidenced in several FDA warning letters, especially in relation to sample handling and reading. These concerns extend to both conventional methods of testing and to rapid methods, methods reliant upon paper documentation and computerized systems, paper documentation (3).
Even with the growing adoption of rapid methods, microbiology laboratories handle a lot of data. Unlike analytical laboratories, however, the data is more often an even mix of quantitative and qualitative data. Examples of data include the result of a settle plate exposed during the environmental monitoring of a cleanroom; the endotoxin content in a sample of water; the weight result from the weighing of a raw material for use in the pharmacopeia Microbial Limits Test and so on.
Despite the current high profile, the subject of data integrity in the pharmaceutical microbiology laboratory has not been afforded very much attention in terms of regulatory guidance. This article looks at two data integrity concerns within the microbiology laboratory, one relating to conventional methods and one to rapid methods, and considers some of the steps that can be taken to address identified weaknesses.
Conventional Methods Environmental monitoring using conventional methods, such as air-sampling, settle plates and contact plates contributes to a cohesive assessment in the assurance of finished product quality. However, traditional culture-based microbiological methods typically possess inherent and unavoidable variability and as such these might result in erroneous conclusions (4). These concerns have been raised in FDA warning letters. Two take two examples: “plate counting, where colony forming units are miscounted” and “missing samples, such as environmental monitoring samples not being taken or dropped on transit to an incubator.” While the latter is careless, colony counting errors can occur where confluent growth occurs.
Some regulators have requested secondary checks for plate reading. This is an erroneous demand and unnecessary – personnel are either good at plate reading or they are not. Solutions to correct colony counting include rapid methods; good training in testing (to ensure that colony numbers fall within the countable range for the plate size); and effective training for reading plates, spotting phenomena like merged colonies, spreading organisms, and diminutive colonies that require reading under magnification and with white light.
Rapid Microbiological Methods There are an array of different rapid microbiological methods, each with their own technologies and testing protocols. Data integrity concerns arise at the design, validation and operation stages. Taking validation, samples need to be representative of what will be tested using the instrument and tested multiple times and by different technicians in order to build in repeatability and robustness. Aspects that give validity to the result, such as limit of detection and limit of quantification (either directly in relation to microorganisms or indirectly through monitoring biological events) need to be introduced.
In terms of operations, data integrity extends to data capture, retention, archiving, and processing. Most rapid methods use computerized systems and here systems should be designed in a way that encourages compliance with the principles of data integrity. Examples include multi-level password control; user access rights which prevent (or audit trail) data amendments; measures to prevent user access to clocks; having automated data capture; ensuring systems have data backup.
When carrying out reviews of rapid microbiological methods five general and important data integrity questions are:
- Is electronic data available?
- Is electronic data reviewed?
- Is meta data (audit trails) reviewed regularly?
- Are there clear segregation of duties?
- Has the system been validated for its intended use?
A satisfactory answer should be available for each of these five key questions.
Summary Data integrity is applicable to microbiology, both conventional and rapid methods, as with every other part of the pharmaceutical operation. This short article has highlighted two areas that microbiologists need to be mindful of in terms of plate counting and computerized systems. Data integrity is an important subject and a regulatory ‘hot topic’; the ideas presented here and wider issues require careful attention by the microbiologist.
References
1. FDA (2016) Data Integrity and Compliance With CGMP, Draft Guidance for Industry, April 2016, U.S. Department of Health and Human Services, Food and Drug Administration, Washington
2. MHRA (2015) MHRA GMP Data Integrity Definitions and Guidance for Industry March 2015, Medicines Healthcare products and Regulatory Agency, London, UK
3. Sandle, T. (2016) Data Integrity Considerations for the Pharmaceutical Microbiology Laboratory, Journal of GXP Compliance, 20 (6): 1-12
4. Tidswell, E. C. and Sandle, T. (2017) Microbiological Test Data - Assuring Data Integrity, PDA Journal of Pharmaceutical Science and Technology, doi:10.5731/pdajpst.2017.008151
 About the author: Dr. Tim Sandle is Head of Microbiology at Bio Products Laboratory, UK and a visiting tutor with the School of Pharmacy and Pharmaceutical Sciences at Manchester University, UK. Tim is a pharmaceutical microbiologist, science writer and journalist with over twenty-five years experience of microbiological research and biopharmaceutical processing. This includes experience of designing, validating and operating a range of microbiological tests including sterility testing, bacterial endotoxin testing, bioburden and microbial enumeration, environmental monitoring, particle counting and water testing.
About the author: Dr. Tim Sandle is Head of Microbiology at Bio Products Laboratory, UK and a visiting tutor with the School of Pharmacy and Pharmaceutical Sciences at Manchester University, UK. Tim is a pharmaceutical microbiologist, science writer and journalist with over twenty-five years experience of microbiological research and biopharmaceutical processing. This includes experience of designing, validating and operating a range of microbiological tests including sterility testing, bacterial endotoxin testing, bioburden and microbial enumeration, environmental monitoring, particle counting and water testing.