As a joint effort, Miacom Diagnostics GmbH and MetaSystems GmbH have developed an ultra-rapid assay that allows 30 minute differentiation of multiple resistant strains (MRSA) and normal, sensitive strains (MSSA) of S. aureus directly from samples without need for sample preparation or costly target amplification.
The assay has now entered the clinical trial phase and will be introduced to the European market to offer a novel solution to physicians to initiate proper antibiotic treatment to patients suffering from a hospital acquired infection such as sepsis or pneumonia. The test doesn´t detect genetic mutations responsible for resistance but instead shows the resistant phenotype of the organism. As this approach is independent of the frequently seen new genetic variations of resistant strains this has advantages over OCR based approach as the outcome of such mutations will always result in the same phenotypic variations.
The new assay will initially be available for use on the Metafer RPI Platform allowing an automated, high-throughput slide analysis directly after the samples are prepared using Miacom’s standard Direct Multiplex Imaging procedure.
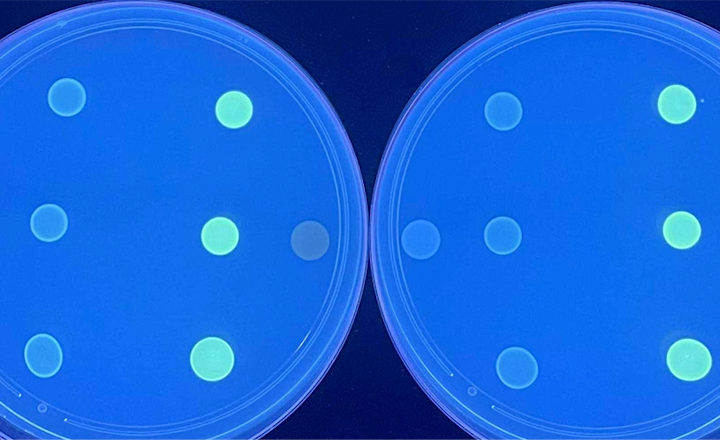
Based on detection of fluorescent light produced by molecular probes when they find their specific target molecules, the MRSA/MSSA differentiation will be available as either a stand-alone assay for screening species of Staphyloccus for MRSA or as a combined multiplex assay detecting more than 10 important Gram positive and Gram negative bacteria in a sepsis. As all of miacom’s assays, this assay relies on the FDA-approved Direct Multiplex Imaging (DMI) technology which allows testing of patient materials directly without the need for amplification in a multiplex fashion differentiating up to 14 pathogens in one single test.
The launch of this addition to Miacom’s existing test kits is expected to take place in 2017.