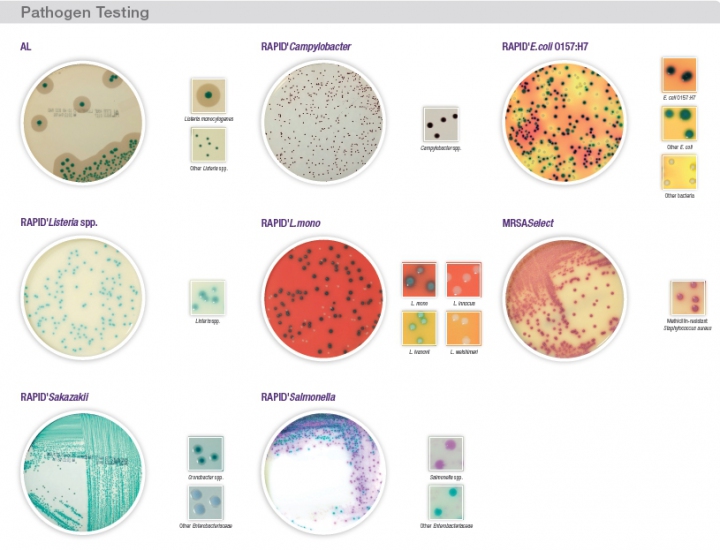
Since Pasteur’s experiments in the 19th century, agar based culture methods have been used to detect and identify microorganisms. Using specific components in their formula, these conventional agar-based culture products allow the determination of a variety of characteristics such as morphology, growth, antibiotic tolerance, metabolic, biochemical, etc. These methods have always been used in microbiological food testing and they remain the gold standard in many ISO protocols and other reference methods.
However, complete bacterial identification still requires up to seven days or more, which is not always suitable with food industry time restraints.
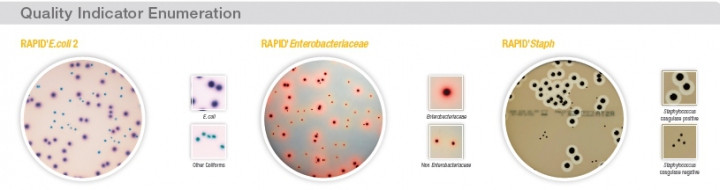
Alternatively, by using chromogenic media, determination of enzymatic activities can be used for bacterial identification. With a variety of synthetic chromogenic substrates included directly in these chromogenic media, specific enzymatic reactions are targeted and microorganisms of interest are directly identified on the agar plate. These alternative chromogenic agar methods allow users to reduce the total time to result with an increase of specificity and ease of use.
The evolution of organic chemistry providing new chromogenic substrate, combined with the expertise in microbiology inherited from the Pasteur background, allow Bio-Rad to offer a complete range of chromogenic media. All Bio-Rad chromogenic methods have been optimized to simplify the workflow and significantly reduce the time to result. By the careful selection of raw materials, these chromogenic media offer a no-compromise quality. All of those qualities have been recognized by the multiple international validations offered: AOAC, NF-Validation, Nordval and Health Canada.
For more information on Bio-Rad complete range of RAPID’Chromogenic media please visit www.bio-rad.com/rapidmedia .
Or watch the video on www.youtube.com/watch