MALDI-TOF MS Rapid Identification for Food Microbiology
Updated: 24th of Feb 2020
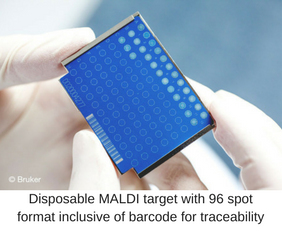
Over the last 30 years the analysis of biomolecules with MALDI time of flight mass spectrometry or MALDI-TOF MS has been well established in academic, pharmaceutical and other industrial laboratories around the world. This success is due to the development of robust benchtop instruments along with dedicated standard protocols and comprehensive libraries for the identification of all kinds of microorganisms including foodborne pathogens.
Thanks to a pretty simple straightforward sample preparation protocol of spotting freshly growing colonies onto a standard steel MALDI target plate, followed by an automated acquisition of a “proteotypic” fingerprint and subsequent bioinformatics pattern profiling, identification of unknown samples is literally done in seconds. Fast and reliable results are essential for any QC lab as it enables them to monitor raw materials and processed food components at critical control points in almost real-time in order to minimize bacterial contamination along the complete food chain and before final product release.
MALDI TOF MS and Microbiology The first MALDI TOF systems were built and used roundabout 1988. A couple of scientists (e.g. K. Tanaka, F. Hillenkamp, M. Karas) had been working on soft ionization techniques in order to create positively charged gas-phase ions of biomolecules after hitting them with a pulsed laser beam. Usually, ablation with such a laser resulted in complete decay of the non-volatile compounds. However, Karas and Hillenkamp developed a technique to co-crystallize the analyte of interest within an UV-absorbing crystal layer of usually hydroxycinnamic acid or derivatives thereof. Since then the term used is matrix-assisted laser desorption/ionization abbreviated to MALDI. Analytical scientists are now able to ionize biomolecules into usually single, positively charged almost intact molecule ions and to focus and accelerate them in a well-defined electrical field. Based on the law of conservation of energy each charged molecule experiences the same energy and can be separated according to its mass in a field free drift region also known as the flight tube. The detector at the end of this tube measures the required time of the different molecules. After a simple calibration with known masses of biomolecules the time of flight (TOF) mass spectrometer can be used as an universal mass analyzer device.
Around 1996 several scientists seperately showed that the analysis of freshly grown microorganisms with MALDI-TOF MS delivers reproducible and typical protein profiles and drew attention to its potential to identify microorganisms in just a few seconds. It took another ten years to develop robust, easy to use benchtop mass spectrometer devices, standardized protocols and most importantly to establish a well-defined, stringently-controlled reference database to match against. Only well characterized type strains, ideally provided from established type culture collections, are used to create the reference database and are cross-checked against field isolates. Finally, discrepancies between the new MALDI-TOF MS and traditional identification methods are eliminated by applying the current gold standard (16S rRNA sequencing).
MALDI-TOF MS and Food Safety Currently, the Bruker reference database contains more than 7000 entries and is continuously growing in agreement with standard QC protocols and aforementioned procedures. In addition, as part of an 'added value' service, the Bruker solution allows us at Biotecon to create customized MALDI reference entries. Any new database update will be tested and validated up-front and finally released only after inspection of all data by at least two experts in the QC department. Although the first systems have solely been used in clinical labs more and more industrial microbiology labs are now using the technique, based on the fact that many typical environmental microorganisms are also well known as food and beverage spoilers and are well represented amongst the current reference entries. The growing number of global users helps to drive further database amendments towards dedicated areas such as the dairy, infant formula and fermented food sectors just to name a few. Interestingly, the first research groups have recently started to create database entries in order to qualify plants, meat, animals, protozoa or other eukaryotes.
Modern MALDI systems utilizing the newest generation of solid state laser technologies are able to create protein profiles nearly every second. With the following bioinformatics pattern profiling running on fast IT processing structures, new identification results are also available each second. In addition, simple straightforward sample preparations and low consumable costs allow more samples to be run at the same time. Ingredients, raw materials, processed food, packaging materials and samples coming in from hygiene control can therefore be controlled in parallel after only a short preparation time. The most convenient sample preparation – the direct colony transfer method for example – allows the preparation of 20 to 30 samples in less than 10 minutes.
In summary MALDI systems deliver fast, accurate identification results translating directly into cost savings, as they help to improve and accelerate the complete food chain controlling process making shorter product cycle rates and smaller warehouse capacities possible. 
Workflow overview illustrates extreme fast time to result due to straightforward, simple sample preparation protocols and subsequent fast MALDI processing.
It has been shown that the described MALDI-TOF MS workflow has significantly shortened turnaround time not only in food safety labs, although in special cases mandatory ISO workflows do not yet allow full implementation of the technique. However, interlaboratory studies and proficiency testing are showing at least the equivalence to currently applied methods and in many cases clearly outperform them. As a consequence ISO working groups are currently evaluating the amendment of current legislation. Meanwhile, MALDI workflows can be directly implemented as an alternative to validation procedures.
Recent efforts in the food and beverage industry clearly show a significant positive impact on identification success after the individual extension of the database with relevant and specific type strains. For example, several QC labs in the brewing sector are now able to correctly identify beer spoilage bacteria including yeasts, to classify the spoilage capacity of certain strains like Lactobacillus brevis and detect wild yeast contaminations. Furthermore, preliminary results from two dairy laboratories are confirming higher positive identification rates and an increased overall scoring level, after adding approximately 300 dairy relevant additional reference entries (Bacillus spp, Cronobcater spp, Campylobacter spp, etc.). Conversely, the number of “no identification”-events have dropped, emphasizing again that continuous monitoring of bacterial contamination and amendment of the screening/identification process (e.g. entering new isolates to the reference library) is a rewarding task.
Conclusion Taken together, modern MALDI-TOF MS systems are delivering fast and reliable identification results within just a few minutes. It has been shown that all available protocols are simple and robust. Manageable workloads and little consumable costs facilitate the application of this method in many laboratories. With more than 7000 entries in the database, the MALDI-TOF MS method provides identification of over 2500 species, including food spoilers, common pathogens, foodborne pathogens, yeasts and molds. The technique additionally allows continuous optimization of the corresponding database and therefore improvements in identification confidence.
This guide was prepared by the product management of BIOTECON Diagnostics MALDI-TOF identification service.
Get the latest updates in Rapid Microbiological Test Methods sent to your email? Subscribe to the free rapidmicrobiology eNewsletter